
Lupine Publishers Group
Lupine Publishers
Menu
Mini Review(ISSN: 2641-6875) 
Whole-Cell Assays for Discovering Novel Efflux Inhibitors for Use as Antibiotic Adjuvants Volume 1 - Issue 2
Ellie Chapman and Sanjib Bhakta*
- Department of Biological Sciences, Institute of Structural and Molecular Biology, University of London, UK
Received:August 30, 2019; Published: September 06, 2019
*Corresponding author: Department of Biological Sciences, Institute of Structural and Molecular Biology, University of London, UK
DOI: 10.32474/CTBM.2019.01.000109
Abstract
Antimicrobial resistance (AMR) is a growing problem worldwide. Resistance to antibiotics can occur in a number of ways, one of which is removal of the drugs from the cell via efflux pump macromolecular machineries. As such, efflux pumps can provide a background level of resistance to many different classes of antimicrobials and are a major contributor to AMR. Inhibition of efflux pumps therefore has the potential to reverse resistance to many antibiotics in one go and is an attractive potential for treating resistant infections. Whilst a number of efflux inhibitors are known, none are currently used clinically due to harmful side effects. Development of novel inhibitors is therefore imperative. The article aims to review accumulation assays and efflux assays, two of the most common laboratory techniques used to identify and characterise candidate efflux inhibitors.
Keywords:Efflux pumps; Efflux inhibitors; Efflux assays; Antimicrobial resistance; Drug discovery
Introduction
Globally, antimicrobial resistance is a rising public health challenge. Particular infections including pneumonia, Tuberculosis (TB), gonorrhoea, and salmonellosis are becoming more difficult to treat. Of new TB cases, 3.5% are either resistant to rifampicin (the most effective first line drug) or are multi-drug resistant, rising to 18% for previously treated individuals [1] Furthermore, there are fears that Neisseria gonorrhoeae has already developed resistance to all currently recommended treatments [2]. There is a desperate need for new antibiotics to treat these most resistant of infections, but the huge costs, long timescale and high attrition rate of drug discovery means that this is a slow process. Twenty classes of antibiotics were discovered between 1940 and 1962, yet only two have been developed since then [3]. Moreover, for any novel antibiotic developed, it is likely that resistance will quickly emerge once it is brought into clinical use, especially with the frequent misuse of antibiotics which drives selection for resistance. Therefore, other strategies must be taken in parallel to antibiotic development, or there will be a continuous arms race of drug development and resulting gain of resistance, a battle we are currently losing.
Figure 1: Schematic representation of the MFS, MATE, SMR, PACE, ABC and RND families of bacterial efflux pumps, plus an outer membrane protein channel (OPM), shown here in a Gram-negative bacterium. RND family efflux pumps comprise of a tripartite complex formed from an inner membrane efflux transporter, an outer membrane channel, and a periplasmic accessory protein. All six families, with the possible exception of the PACE family, also have representatives in both Grampositive and acid-fast bacteria. Bold arrows indicate the direction of drug efflux, and dashed arrows show ion movement.
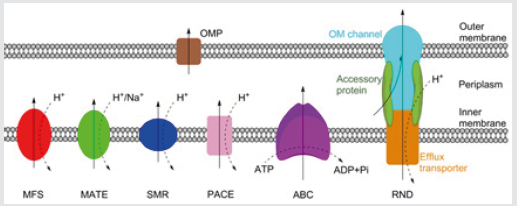
Antibiotic resistance can occur via acquired or intrinsic mechanisms. Acquired resistance, typically via horizontal transfer or spontaneous mutation, often functions by altering the drug target or production of enzymes which degrade the antibiotic. Acquired resistance, gained in response to antibiotic treatment, is usually only effective against a single drug. Intrinsic resistance, on the other hand, refers to the non-specific mechanisms of antibiotic resistance evolved ancestrally, including the impermeable outer membrane of Gram-negative or acid-fast group of bacteria, and drug efflux pumps which remove drugs from the cell [4]. There are currently six families of bacterial efflux pumps identified: the ATPBinding Cassette (ABC) family, the Major Facilitator Superfamily (MFS), the Multidrug And Toxin Extrusion (MATE) family, the Small Multidrug Resistance (SMR) family, the Resistance-Nodulation-Cell Division (RND) superfamily and the Proteobacterial Antimicrobial Compound Efflux (PACE) family, which has not yet been structurally characterised. The ABC family hydrolyse ATP directly to drive efflux, whereas the other five utilise transmembrane ion gradients [5]. Whereas the RND family directly effluxes antibiotics across both membranes, the other five families only transport antibiotics across the inner membrane. From the periplasm, drugs can exit the cell via outer membrane protein channels or by entering the RND complex (Figure 1).
Efflux pumps are often non-specific, and as such can provide resistance to a wide range of antimicrobials. They have been implicated in contributing towards the multi-drug resistant phenotypes of Mycobacterium tuberculosis [6], Pseudomonas aeruginosa [7], Neisseria gonorrhoeae [8], and Streptococcus pneumoniae [9], amongst others. Inhibition of drug efflux is therefore an exciting prospect for treating drug resistant bacteria and may enable old antibiotics to re-enter clinical usage. There is compelling evidence that the use of efflux pump inhibitors as an adjuvant may aid treatment of resistant infections of many types [6-12]. However, despite a number of potent efflux inhibitors being known, none have entered clinical use. In most cases this is because the compounds are toxic at the concentrations required to inhibit efflux [13]. There is therefore a pressing need to develop novel clinical efflux inhibitors. To achieve this, assays are needed to validate the inhibitory activity of novel compounds. One way this can be achieved is by using standard antibiotic susceptibility testing, such as the resazurin-based microplate assay to determine if the putative inhibitor, at sub-MIC concentrations, is able to lower the MIC of a known antibiotic. This method has the benefit of being relatively easy and high-throughput; furthermore, it is possible to combine this method with mutants of efflux pumps to confirm that the effect on the MIC is occurring specifically via inhibiting efflux, and even to identify which efflux pump is inhibited [14]. However, using reduction of MICs to identify and validate efflux inhibitors is fairly insensitive, and so is of limited use. Only large changes to efflux will likely have an effect on MICs, and so less potent inhibitors may be dropped out. Furthermore, as this method does not measure efflux, it is difficult to directly attribute changes in MIC to efflux inhibition [15]
A more direct way is therefore needed to study the effect of candidate inhibitors on efflux. One way is to follow the movement of an efflux pump substrate, often a fluorescent molecule, into and out of bacterial cells, and use this as a measure of efflux activity. Many different molecules are used to measure efflux, with ethidium bromide and Nile red being two of the most common. Ethidium bromide fluoresces strongly when bound to DNA, and Nile red fluoresces when in non-polar environments such as the membrane [16,17]. This therefore gives these molecules the advantage that they fluoresce differentially in extra- and intracellular environments, providing a sensitive indication of rate of efflux from the cell, and helping eliminate background fluorescence. These methods fall into two main categories; those which follow the accumulation of the molecule within the cell, and those which follow its efflux.
Accumulation Assays
Whilst there are variations, most accumulation assays typically follow a similar procedure. At the start of the assay, there is no dye added to the bacteria. This is then added to the reaction, and its accumulation within the cells followed over time, typically by measuring the fluorescence with dyes such as ethidium bromide. Eventually, accumulation will tail off, with fluorescence reaching a steady state. This reflects an equilibrium being achieved between influx and efflux of the dye. This assay can be performed with added efflux inhibitors [18]. By inhibiting efflux, more dye accumulates within the cells compared to untreated ones, with steady state being achieved at a higher fluorescence. This assay can therefore be used as a very simple test to validate the inhibitory activity of a candidate efflux inhibitor [19]. Similarly, accumulation assays are often used to observe changes in efflux ability in knockout, knockdown or overexpression mutants.
If a knockout/knockdown mutant accumulates more dye, it can be assumed that the gene encoded a protein important for drug efflux, or a regulator of these, and vice versa with overexpression mutants. These two approaches can be combined, with different mutants treated with efflux inhibitors to see if they have a greater or lesser effect on dye accumulation than for wild-type cells. This can help determine which efflux pump the inhibitor affects [7]. However, there are problems with using accumulation assays, the most important being that accumulation is not a direct measure of efflux. Rather, it reflects a number of factors, predominantly the balance of influx and efflux rates. Influx depends greatly on the permeability of bacterial membranes, which can vary greatly between even closely related strains due to differing membrane compositions [20]. Therefore, unless influx rates are known, kinetic data cannot be obtained from accumulation assays and results remain qualitative. Whilst this limits usage of accumulation assays to comparisons between isogenic mutants, or groups treated with different inhibitors, the assay remains a conclusive way to determine if a molecule possesses inhibitory activity, and so is frequently used to validate new efflux inhibitors.
Efflux Assays
If a quantitative measure of efflux is required, then a more direct efflux assay should be used. This follows a similar premise to accumulation assays, but instead involves preloading the cells with dye and following its subsequent efflux. To achieve this, cells are incubated with a dye or other efflux pump substrate, and a known efflux inhibitor such as CCCP. This causes the dye to accumulate to a maximum level. Then, the cells are washed to remove the inhibitor and any remaining extracellular dye. The cells are then reenergised, typically with glucose, which restarts efflux. The movement of the dye out of the cells can be followed by recording the decreasing fluorescence [15]. As this method is a direct measure of efflux, kinetic data can be obtained for efflux rates, which allows comparisons to be made more broadly, rather than just between isogenic species. In much the same way as with accumulation assays, modifications can be made to study the effects of putative inhibitors or different mutations on efflux rates [12,21].
Efflux assays are very sensitive, and they allow for validation and characterisation of novel inhibitors, which may potentially have clinical usage. Whilst the efflux assay is widely used, it is not always applicable. Non-fermenter bacteria, including Pseudomonas and Acinetobacter, are unable to metabolise glucose, and so cannot be easily reenergised. This means that efflux assays can be unsuitable for some bacteria, and instead accumulation assays are more commonly used [7,22].
Limitations with these Assays
A fundamental problem with both types of assay is that using ethidium bromide or another dye to measure efflux or accumulation is of limited clinical relevance, and may not reflect well the efflux of any particular antibiotic. This can be due to the dye and antibiotic having very different kinetics of efflux, and furthermore, they may not even be substrates for the same efflux pumps. In addition, as ethidium bromide intercalates with DNA, there is a lag time in efflux in which it dissociates, followed potentially by a two-step efflux mechanism in which it is first transported to the periplasm. This can lead to underestimates of efflux rate, and so may be a poor reflection of efflux rates of antibiotics [23]. Therefore, where possible, it is better to use the antibiotic of interest itself as a direct measure of efflux, although this tends to be far more difficult experimentally. Certain antibiotics, such as fluroquinolones and tetracyclines have endogenous fluorescence which enables their accumulation to be followed [24]. For non-fluorescent antibiotics, Mass-Spectroscopy (MS) can be used to directly study their accumulation. A recent proposed joint protocol for spectrofluorimetric and MS analyses suggests that the two methods are complementary and together can accurately measure antibiotic accumulation, demonstrated with fluroquinolones [25]. MS analyses, rather than spectrofluorometric, may also provide a better way to screen natural compounds for efflux inhibitory activity. Many natural compounds have endogenous fluorescence, which can make it hard to isolate and interpret fluorescence changes due to dye accumulation or efflux. As before, the actual antibiotic, rather than a dye, could be used, and MS used to determine how much accumulates with and without the candidate inhibitor.
One of the biggest problems facing the development of novel efflux inhibitors is the lack of high-throughput assays to validate putative compounds. Whilst both the accumulation and efflux assays are relatively easy to perform and can reliably confirm if inhibition occurs, both are limited on throughput. Therefore, whilst some in silico screening has been performed [26], limitations in throughput have so far prevented large-scale screening of libraries in vitro. Instead, the search for novel inhibitors has relied extensively on prior knowledge to select candidates for validation. Whilst the hit rate with this has been relatively high, the overall number of new inhibitors found has been low, and it is rare to identify completely novel inhibitors in this way. This is in part why no inhibitors have made their way into clinical usage, as many are closely related and as such are similarly toxic. Development of high-throughput screening assays for novel inhibitors is therefore necessary if efflux inhibitors are to progress clinically. Recently, the Back assay was developed, which uses a 96-well plate format combined with MS. This was able to test in triplicate 12 compounds at 4 concentrations each, for two different Escherichia coli strains [27]. This progression to more high-throughput screening is likely to be the driving force behind development of novel efflux inhibitors, and further work needs to be done to optimise assays before large scale-screening of compound libraries can be performed. Ultimately, the development of clinical efflux inhibitors used therapeutically as antibiotic adjuvants may be what turns the tide in the battle against antibiotic resistance.
Acknowledgment
The authors would like to thank the British Society for Antimicrobial Chemotherapy, without whose funding this work would not have been possible. We also wish to thank Dr Arundhati Maitra for her time and advice when writing the article, as well as for help with ChemBioDraw.
References
- World Health Organization (2018) Global Tuberculosis Report 2018.
- Unemo M, Golparian D, Eyre DW (2019) Antimicrobial Resistance in Neisseria gonorrhoeae and Treatment of Gonorrhea. Methods Mol Biol 37-58.
- Mantravadi P, Kalesh K, Dobson R, Hudson A, Parthasarathy A (2019) The Quest for Novel Antimicrobial Compounds: Emerging Trends in Research, Development, and Technologies. Antibiotics (Basel) 24: 8(1).
- Admassie M (2018) Current Review on Molecular and Phenotypic Mechanism of Bacterial Resistance to Antibiotic. Science PG 7(2): 13-19.
- Du D, Wang Kan X, Neuberger A, van Veen HW, Pos KM, et al. (2018) Multidrug efflux pumps: Structure, function and regulation. Nat Rev Microbiol 16(9): 523-539.
- Pule CM, Sampson SL, Warren RM, Black PA, van Helden PD (2016) Efflux pump inhibitors: Targeting mycobacterial efflux systems to enhance TB therapy. J Antimicrob Chemother 71(1): 17-26.
- Siriyong T, Srimanote P, Chusri S, Yingyongnarongkul B, Suaisom C (2017) Conessine as a novel inhibitor of multidrug efflux pump systems in Pseudomonas aeruginosa. BMC Complement Altern Med 17(1): 405.
- Golparian D, Shafer WM, OhnishiM, Unemo M (2014) Importance of Multidrug Efflux Pumps in the Antimicrobial Resistance Property of Clinical Multidrug-Resistant Isolates of Neisseria gonorrhoeae. Antimicrob Agents Chemother 58(6): 3556-3559.
- Iannelli F, Santoro F, Santagati M, Docquier JD, Lazzeri E, et al. (2018) Type M Resistance to Macrolides Is Due to a Two-Gene Efflux Transport System of the ATP-Binding Cassette (ABC) Superfamily. Front Microbiol 31(9): 1670.
- Shiu WKP, Malkinson JP, RahmanMM, Curry J, Stapleton P, et al. (2013) A new plant-derived antibacterial is an inhibitor of efflux pumps in Staphylococcus aureus. Int J Antimicrob Agents 42(6): 513-518.
- Kakarla P, Floyd J, Mukherjee M, Devireddy AR, Inupakutika MA, et al. (2017) Inhibition of the multidrug efflux pump LmrS from Staphylococcus aureus by cumin spice Cuminum cyminum. Arch Microbiol 199(3): 465-474.
- Machado D, Fernandes L, Costa SS, Cannalire R, Manfroni G (2017) Mode of action of the 2-phenylquinoline efflux inhibitor PQQ4R against Escherichia coli. Peer J 26(5): 3168.
- Mahmood HY, Jamshidi S, Sutton JM, Rahman KM (2016) Current Advances in Developing Inhibitors of Bacterial Multidrug Efflux Pumps. Curr Med Chem 23(10): 1062-1081.
- Balganesh M, Dinesh N, Sharma S, Kuruppath S, Nair AV (2012) Efflux pumps of Mycobacterium tuberculosis play a significant role in antituberculosis activity of potential drug candidates. Antimicrob Agents Chemother 56(5): 2643-2651.
- Blair JMA, Piddock LJV (2016) How to Measure Export via Bacterial Multidrug Resistance Efflux Pumps. MBio 7.
- Olmsted J, Kearns DR (1977) Mechanism of ethidium bromide fluorescence enhancement on binding to nucleic acids. Biochemistry 16(16): 3647-3654.
- Greenspan P, Fowler SD (1985) Spectrofluorometric studies of the lipid probe, nile red. J Lipid Res 26(7): 781-789.
- Rodrigues L, Viveiros M, Ainsa JA (2015) Measuring Efflux and Permeability in Mycobacteria. Methods Mol Biol 1285: 227-239.
- Danquah CA, Kakagianni E, Khondkar P, Maitra A, Rahman M, et al. (2018) Analogues of Disulfides from Allium stipitatum Demonstrate Potent Anti-tubercular Activities through Drug Efflux Pump and Biofilm Inhibition. Sci Rep 8(1): 1150.
- Sohlenkamp C, Geiger O (2016) Bacterial membrane lipids: diversity in structures and pathways. FEMS Microbiol Rev 40(1): 133-159.
- Smith HE, Blair JMA (2014) Redundancy in the periplasmic adaptor proteins AcrA and AcrE provides resilience and an ability to export substrates of multidrug efflux. J Antimicrob Chemother 69(4): 982-987.
- Richmond GE, Chua KL, Piddock LJV (2013) Efflux in Acinetobacter baumannii can be determined by measuring accumulation of H33342 (bis-benzamide). J Antimicrob Chemother 68(7): 1594-1600.
- Bohnert JA, Karamian B, Nikaido H (2010) Optimized Nile Red efflux assay of AcrAB-TolC multidrug efflux system shows competition between substrates. Antimicrob Agents Chemother 54(9): 3770-3775.
- Vergalli J, Dumont E, Cinquin B, Maigre L, Pajovic J (2017) Fluoroquinolone structure and translocation flux across bacterial membrane. Sci Rep 7: 9821.
- Dumont E, Vergalli J, Conraux L, Taillier C, Vassort A (2018) Antibiotics and efflux: Combined spectrofluorimetry and mass spectrometry to evaluate the involvement of concentration and efflux activity in antibiotic intracellular accumulation. J Antimicrob Chemother 74(1): 58-65.
- Choudhury D, Talukdar A, Chetia P, Bhattacharjee A, Choudhury M (2016) Screening of Natural Products and Derivatives for the Identification of RND Efflux Pump Inhibitors. Comb Chem High Throughput Screen 19(9): 705-713.
- Widya M, Pasutti WD, Sachdeva M, Simmons RL, Tamrakar P (2019) Development and Optimization of a Higher-Throughput Bacterial Compound Accumulation Assay. ACS Infect Dis 5(3): 394-405.

Top Editors
-

Mark E Smith
Bio chemistry
University of Texas Medical Branch, USA -

Lawrence A Presley
Department of Criminal Justice
Liberty University, USA -

Thomas W Miller
Department of Psychiatry
University of Kentucky, USA -

Gjumrakch Aliev
Department of Medicine
Gally International Biomedical Research & Consulting LLC, USA -

Christopher Bryant
Department of Urbanisation and Agricultural
Montreal university, USA -

Robert William Frare
Oral & Maxillofacial Pathology
New York University, USA -

Rudolph Modesto Navari
Gastroenterology and Hepatology
University of Alabama, UK -

Andrew Hague
Department of Medicine
Universities of Bradford, UK -

George Gregory Buttigieg
Maltese College of Obstetrics and Gynaecology, Europe -

Chen-Hsiung Yeh
Oncology
Circulogene Theranostics, England -
.png)
Emilio Bucio-Carrillo
Radiation Chemistry
National University of Mexico, USA -
.jpg)
Casey J Grenier
Analytical Chemistry
Wentworth Institute of Technology, USA -
Hany Atalah
Minimally Invasive Surgery
Mercer University school of Medicine, USA -

Abu-Hussein Muhamad
Pediatric Dentistry
University of Athens , Greece

The annual scholar awards from Lupine Publishers honor a selected number Read More...





