Lupine Publishers Group
Lupine Publishers
Menu
ISSN: 2637-4706
Mini Review(ISSN: 2637-4706) 
Development of Small Molecule Inhibitor Targeting Aminoacyl-Trna Synthetase Interacting Multifunctional Protein 2 (AIMP2)-DX2 As Anti-Cancer Therapeutics Volume 3 - Issue 5
Seungbeom Lee*
- Department of Chemistry and Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, California, United States
Received: December 17, 2020; Published: February 19, 2021
Corresponding author: Seungbeom Lee, Department of Chemistry and Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, California 92037, United States
DOI: 10.32474/DDIPIJ.2021.03.000172
Abstract
Recently, noncanonical roles of Multi-tRNA Synthetase Complex (MSC) proteins including Aminoacyl-tRNA synthetase Interacting Multifunctional Proteins (AIMPs) have been revealed and served as a chance to develop novel therapeutics with their unique mechanisms. The disease specific splicing variant of AIMP2 (AIMP2-DX2) has recently been considered a novel promising therapeutic target in several cancers. However, few studies on developing small molecule inhibitors of AIMP2-DX2 have been reported. In this paper, the brief summary for AIMP2-DX2 as a medicinal chemistry target, has been demonstrated.
Keywords: Drug Discovery; Medicinal Chemistry; AIMP2-DX2; Anti-cancer; Alternative Splicing Variant
Mini Review
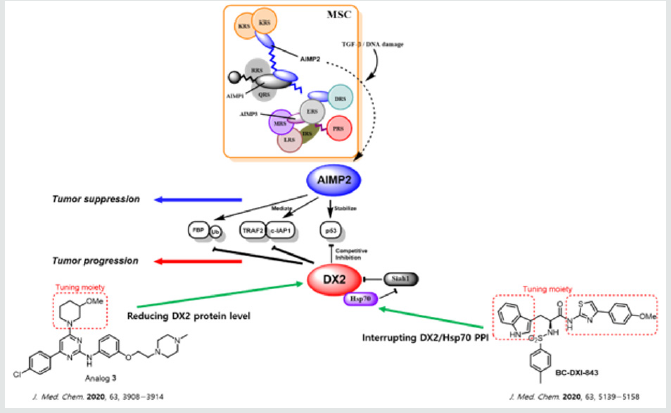
In human, Multi-tRNA Synthetase Complex (MSC) which is composed nine kinds of Aminoacyl-tRNA Synthetases (ARSs) and three kinds of ARS Interacting Multifunctional Proteins (AIMPs), plays an essential role in synthesizing aminoacyl-tRNA with essential amino acids. AIMPs have auxiliary functions to assemble and integrate MSC components [1,2]. Recently, since the noncanonical roles of many elements of MSC have been revealed, ARSs and AIMPs have been disclosed as prospective therapeutic targets for several critical diseases [3-7]. In proper conditions, AIMP2 is released from MSC to cytoplasm and plays an important role in tumor suppressing process via TNF-α, TGF-β, and p53 pathways (Figure 1) [8-11]. Contrastively, the disease specific splicing variant of AIMP2 (AIMP2-DX2, also called DX2) which exon 2 of AIMP2 full length consisting of 4 exons is excised, inhibits these tumor suppressing activities of AIMP2. The expression level of DX2 has been higher in chemoresistant ovarian cancers, prostate, nasopharyngeal and especially lung cancers than that of normal cells. Thus, DX2 is served as the oncogenic target without adverse effect even when it was fully deleted, supported by several DX2 knock-out experiments [12-16]. In 2013, the First Small Molecule Inhibitor (BC-DXI01) identified via high-throughput screening of the synthetic chemical library, has been reported by Kim and co-workers [17]. This screening covered the massive structural diversity due to the library size including 2231 small molecules and various scaffolds. Recently, in 2016, the other DX2 inhibitor (SLCB050) which exhibited inhibitory activity on DX2-p14/ARF interaction, has been reported by Park and co-workers [18]. However, the early inhibitor BC-DXI01 and SLCB050 were unsatisfactory in terms of potency and applicable spectrum of tumor cells. BC-DXI01 exhibited moderate to low inhibitory activity on DX2-luciferase (IC50 = 20.1 μM) and SLCB050 exhibited poor cytotoxic activity on NSCLC (GI50 > 50 μM) [17,18]. More recently, in 2020, two advanced medicinal chemistry works have been published (Figure 1) [19,20]. Analog 3 which has been reported by Suh and co-workers, exhibited 3.58 μM IC50 value of DX2-luciferase and significant inhibitory activity on H460 cells (GI50 = 0.60 μM). Analog 3 was optimized from HTS hit analog 1 through synthesizing analogs with 3-parts substitution of the center 2-aminopyrimidine moiety as a strategy. Structure- Activity Relationship (SAR) of analogs indicated that methyl group of analog 1 as A part substitution moiety is the tunable site which is finally optimized to 3-methoxypiperidine moiety in analog 3. The cytotoxic activities of analog 3 on several cell lines revealed the corelation between GI50 and the expression level of DX2 protein of each cell lines. According to further in vitro experiments, analog 3 inhibited DX2 protein expression but exhibited no inhibition of the level of DX2 mRNA. These results implicated that the mechanism of action of analog 3 is related to degradation or translation of DX2 but not transcription of DX2. In mice xenograft model, analog 3 inhibited tumor growth without notable toxic symptom [19]. Level of DX2 in cells is regulated by Siah1 protein which decrease level of DX2 via ubiquitination-degradation sequence, however, molecular chaperone protein Hsp70 binds to DX2 and interferes ubiquitination of Siah1. Thus, inhibition of the Protein-Protein Interaction (PPI) between DX2 and Hsp70 enables to lead to anticancer effect [21]. Lee and co-workers have disclosed sulfonamidebased PPI inhibitor (BC-DXI-843) optimized from HTS hit of DX2 (BC-DXI-04). As the chiral sulfonamide moiety at α-position of amide was the standard scaffold, 3 parts of side chains were modified and optimized. BC-DXI-843 exhibited higher inhibitory activity on DX2 with sub-micromolar DX2 IC50 (0.92 μM) and low-micromolar GI50 (1.20 μM) against A549 compared to that on AIMP2 and WI- 26 cells (IC50 and GI50 >100 μM) as well as significant inhibitory activity on tumor of mice xenograft model [20]. Through examples of these advanced small molecular DX2 inhibitors, we suggest that there are some challenges such as selectivity, physicochemical properties, chronic or acute toxicity and PK issue to develop further DX2 inhibitors. Particularly, the selectivity between AIMP2 is able to be a critical point to develop further DX2 inhibitors since DX2 derived from AIMP2 is suspected of its similarity to AIMP2 which is the essential tumor suppressor. The key strategy must be tightly checking toxicities as well as improving potency and PK profiles with the proper selectivity to DX2 versus AIMP2.
References
- Han JM, Lee MJ, Park SG, Lee SH, Razin E (2006) Hierarchical network between the components of the multitRNA synthetase complex: Implications for complex formation. J Biol Chem 281(50): 38663-38667.
- Park SG, Choi EC, Kim S (2010) Aminoacyl-tRNA synthetase interacting multifunctional proteins (AIMPs): A triad for cellular homeostasis. IUBMB Life 62(4):296-302.
- Guo M, Schimmel P (2013) Essential nontranslational functions of tRNA synthetases. Nat Chem Biol 9(3): 145-153.
- Jo SM, Kim Y, Jeong YS, Hee Oh Y, Park K, et al. (2013) Rapid detection of exon 2-deleted AIMP2 mutation as a potential biomarker for lung cancer by molecular beacons. Biosens Bioelectron 15(46): 142-149.
- Jung J Y, Kim E Y, Kim A, Chang J, et al. (2017) Ratio of autoantibodies of tumor suppressor AIMP2 and its oncogenic variant is associated with clinical outcome in lung cancer. J Cancer 8(8): 1347−1354.
- Kwon N H, Fox P L, Kim S.(2019) Aminoacyl-tRNA synthetases as therapeutic targets. Nat Rev Drug Discovery 18(8): 629-650.
- Kim M, Kim H, Kim D, Park C, Huh Y, et al. (2019) Fluorescence-based analysis of noncanonical functions of aminoacyl-tRNA synthetase-interacting multifunctional proteins (AIMPs) in peripheral nerves. Materials 12(7): 1064.
- Kim MJ, Park BJ, Kang YS, Kim HJ, Park JH, et al. (2003) Downregulation of FUSE-binding protein and c-myc by tRNA synthetase cofactor p38 is required for lung cell differentiation. Nat Genet 34(3): 330-336.
- Han JM, Park BJ, Park SG, Oh YS, Choi SJ, et al. (2008) AIMP2/p38, the scaffold for the multi-tRNA synthetase complex,responds to genotoxic stresses via p53. PNAS 105(32): 11206-11211.
- Choi JW, Kim DG, Park MC, Um JY, Han JM, et al. (2009) AIMP2 promotes TNFalpha-dependent apoptosis via ubiquitin-mediated degradation of TRAF2. JCell Sci 122(pt15): 2710-2715.
- Kim S, You S, Hwang D (2011) Aminoacyl-tRNA synthetases and tumorigenesis: More than housekeeping. Nat Rev Cancer 11(10): 708-718.
- Choi JW, Kim DG, Lee AE, Kim HR, Lee JY, et al. (2011) Cancer-associated splicing variant of tumor suppressor AIMP2/p38: Pathological implication in tumorigenesis. PLoS Genet 7(3): 1001351.
- Choi JW, Lee JW, Kim JK, Jeon HK, Choi JJ, et al. (2012) Splicing variant of AIMP2 as an effective target against chemoresistant ovarian cancer. J Mol Cell Biol 4(3): 164-173.
- Won YS, Lee SW (2012) Selective regression of cancer cells expressing a splicing variant of AIMP2 through targeted RNA replacement by trans-splicing ribozyme. J Biotechnol 158(1-2): 44-9.
- Chang SH, Chung YS, Hwang SK, Kwon JT, MinaiTehrani A (2012) Lentiviral vector-mediated shRNA against AIMP2-DX2 suppresses lung cancer cell growth through blocking glucose uptake. Mol Cells 33(6): 553-562.
- Cao Q, Zhang J, Zhang T (2018) AIMP2-DX2 promotes the proliferation, migration, and invasion of nasopharyngeal carcinoma cells. BioMed Res Int 2018: 9253036.
- Oh AY, Jung YS, Kim J, Lee JH, Cho JH, et al. (2016) Inhibiting DX2-p14/ARF interaction exerts antitumor effects in lung cancer and delays tumor progression. Cancer Res 76(16): 4791-4804.
- Lee HS, Kim DG, Oh YS, Kwon NH, Lee JY, et al. (2013) Chemical suppression of an oncogenic splicing variant of AIMP2 induces tumour regression. Biochem 454(3): 411-416.
- Lee S, Kim DG, Kim K, Kim T, Lim S, et al. (2020) 2-Aminophenylpyrimidines as Novel Inhibitors of Aminoacyl-tRNA Synthetase Interacting Multifunctional Protein 2 (AIMP2)-DX2 for Lung Cancer Treatment. JMed. Chem 63(8): 3908-3914.
- Sivaraman A, Kim DG, Bhattarai D, Kim M, Lee HY, et al. (2020) Synthesis and Structure–Activity Relationships of Arylsulfonamides as AIMP2-DX2 Inhibitors for the Development of a Novel Anticancer Therapy. Jof Med Chem 63(10): 5139-5158.
- Lim S, Cho HY, Kim DG, Roh Y, Son S, et al. (2020) Targeting the interaction of AIMP2-DX2 with HSP70 suppresses cancer development. Nat Chem Biol 16(1) 31-41.

Top Editors
-

Mark E Smith
Bio chemistry
University of Texas Medical Branch, USA -

Lawrence A Presley
Department of Criminal Justice
Liberty University, USA -

Thomas W Miller
Department of Psychiatry
University of Kentucky, USA -

Gjumrakch Aliev
Department of Medicine
Gally International Biomedical Research & Consulting LLC, USA -

Christopher Bryant
Department of Urbanisation and Agricultural
Montreal university, USA -

Robert William Frare
Oral & Maxillofacial Pathology
New York University, USA -

Rudolph Modesto Navari
Gastroenterology and Hepatology
University of Alabama, UK -

Andrew Hague
Department of Medicine
Universities of Bradford, UK -

George Gregory Buttigieg
Maltese College of Obstetrics and Gynaecology, Europe -

Chen-Hsiung Yeh
Oncology
Circulogene Theranostics, England -
.png)
Emilio Bucio-Carrillo
Radiation Chemistry
National University of Mexico, USA -
.jpg)
Casey J Grenier
Analytical Chemistry
Wentworth Institute of Technology, USA -
Hany Atalah
Minimally Invasive Surgery
Mercer University school of Medicine, USA -

Abu-Hussein Muhamad
Pediatric Dentistry
University of Athens , Greece

The annual scholar awards from Lupine Publishers honor a selected number Read More...