Lupine Publishers Group
Lupine Publishers
Menu
Review Article(ISSN: 2641-6875) 
Analysis of the Microbial Structure in Daqu for the Main Types of Chinese Baijiu Volume 1 - Issue 1
XingLin Han1,2, DeLiang Wang2, JinYuan Sun1, WuJiu Zhang2 and BaoGuo Sun1*
- 1Beijing Laboratory for Food Quality and Safety, Beijing Technology & Business University (BTBU), China
- 2China National Research Institute of Food and Fermentation Industries, China
Received: January 21, 2021; Published: February 07, 2022
*Corresponding author: BaoGuo Sun, Beijing Laboratory for Food Quality and Safety, Beijing Technology & Business University (BTBU), China
DOI: 10.32474/CTBM.2022.02.000146
Abstract
This investigation undertook a systematic analysis of the microbial structure of six kinds of daqu from different types of Chinese baijiu. From the results, it was noted that the prokaryotic microbial species present in daqu were much more complex than the eukaryotic microorganisms, but the number of eukaryotic microorganisms was greater than that of prokaryotic microbes. The investigation identified 8 phyla, 74 classes, 130 orders, 203 families, and 255 genera for eukaryotic microbes and 11 phyla, 74 classes, 116 orders, 411 families, and 473 genera for prokaryotic microorganisms. The main eukaryotic microbe in daqu was Ascomycota, followed by Basidiomycota; Ascomycota species are mainly moulds and yeasts, and they play a major role in of baijiu fermentation. The main prokaryotic microorganism in daqu was Firmicutes, followed by Actinobacteria, Proteobacteria, Bacteroidetes, Euryarchaeota and Planctomycetaceae. The number of eukaryotic microbes OTUs were ranked in the order Nongxiang> Zhima-xiang>Qing-xiang>Feng-xiang>Chi-xiang>Jiang-xiang, and the abundance values were ranked in the order Qingxiang> Nong-xiang>Jiang-xiang>Zhima-xiang>Chi-xiang. The number of prokaryotic microorganisms OTUs were ranked in the order Qing-xiang>Nong-xiang>Zhima-xiang>Feng-xiang>Chi-xiang>Jiang-xiang, and the abundance values were ranked in the order Qing-xiang>Nong-xiang>Zhima-xiang>Feng-xiang>Chi-xiang>Jiang-xiang.
Keywords: Chinese Baijiu; High-Throughput Sequencing; Lactobacillaceae; Eurotium
Introduction
Chinese baijiu is one of the world’s oldest distilled alcoholic beverages and is very popular in China. It uses daqu as the fermentation starter, which is a saccharifying and fermenting agent that has a significant impact on the flavour of the product [1]. However, the daqu is different when used in baijiu brewing with different technologies. For example, Jiang-xiang baijiu uses a daqu with a higher temperature in the fermentation process, but Qing-xiang baijiu uses a daqu with a lower temperature in the fermentation process [2]. Moreover, the different kinds of daqu use different raw materials. Therefore, the microbial structure should be different for the daqu for the different types of baijiu. The microorganisms found in daqu have been analysed and reported in the literature; for example, in 2007, Yang Daiyong et al. used the selective culture method to carry out a pilot study of the microbial composition in daqu from Maotai [3]. In 2010, Meng Zhen et al. used PCR-DGGE technology to analyse the bacterial community structure of daqu and found that Firmicutes, Lactobacillaceae, Bacillaceae and Thermoactinomycetaceae were the main bacteria [4]. In 2020, Chen meng-en et al. used the selective culture method to identify Pantoea, Bacillus, Leuconostoc, Thermoascus, Aspergillus, and Thermomyces in Tao-rong daqu [5]. In 2020, Sun li-lin et al. used high-throughput sequencing technology to identify Oceanobacillus, Bacillales, Aspergillus, Thermoascus and Thermomyces in Jiang-xiang daqu [6]. However, this previous research perhaps involved only one kind of daqu and perhaps used the traditional culture method, yielding a large error value [7-10]. High throughput sequencing simultaneously sequences several million to billion DNA molecules and allows in-depth, meticulous, and global analysis; hence, it is called deep sequencing. It enables the rapid and convenient determination of the complicated microbial structure of a sample. The analytical ability and precision of PCR-DGGE technology is markedly more advanced than those of 16S rDNA sequence homology technology, which was the first method of analysing complex multi-strain samples and, at present, has not been used for analysing the microbial structure of daqu from different types of Chinese baijiu [11-14]. This study used high-throughput sequencing technology to research and analyse microorganisms in the main type of baijiu of Nong-xiang daqu, Qing-xiang daqu, Jiang-xiang daqu, Zhima-xiang daqu, Chi-xiang daqu and Feng-xiang daqu.
Materials and Methods
Experimental Samples
Ten typical daqu samples of Nong-xiang, Qing-xiang, Jiangxiang, Zhima-xiang, Chi-xiang and Feng-xiang were used. Each sample was broken down and mixed uniformly to obtain a mixed sample representing Nong-xiang daqu (ND), Qing-xiang daqu (QD), Jiang-xiang daqu (JD), Zhima-xiang daqu (ZD), Chi-xiang daqu (CD) and Feng-xiang daqu (FD), and microbial analysis was performed.
Testing Methods and Instruments
To analyse the microbial structure of daqu, Illumina MiSeq sequencing technology was used. The samples were mixed and well distributed and were analysed by the China National Research Institute of Food and Fermentation Industries.
Results and Analysis
Microbial Structure from All the Daqu Types (1,5,15-17)
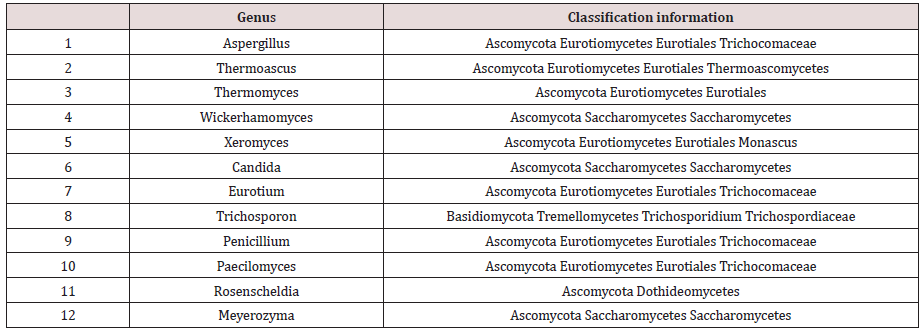
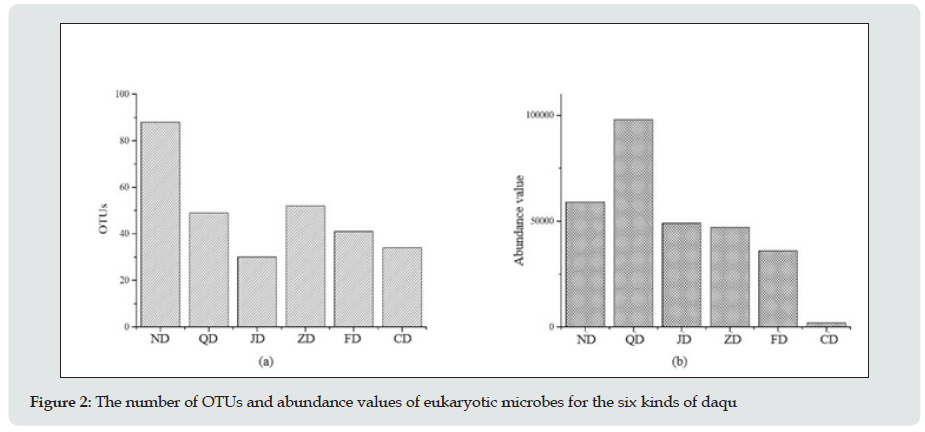
To determine the microbial diversity in daqu samples, UCLUST software was used for the cluster analysis of the orders. Ninetyseven percent of the orders were similar, and UCLUST was used to designate operational taxonomic units (OTUs). OTUs indicate a group of orders that came from the same classified unit. The results regarding the kinds and numbers of microbes from daqu of 6 different types of baijiu are shown in Figure 1. Prokaryotic microorganisms accounted for 73% of the total kinds, but they accounted for only 38% of the total number. High-throughput sequencing revealed 8 phyla, 74 classes, 130 orders, 203 families, and 255 genera of eukaryotic microbes and 11 phyla, 74 classes, 116 orders, 411 families, and 473 genera of prokaryotic microorganisms. (Table 1) shows the eukaryotic microbes whose abundance value was more than 0.5%, which were detected in this experiment. From the statistical analysis results, the main eukaryotic microbe in daqu was Ascomycota, followed by Basidiomycota; therein, Ascomycota are mainly mould and yeast, and they play a major role in baijiu fermentation. Additionally, Wallemia, Trichosporon and Meyerozyma were not reported in the previous literature. Wallemia is mainly spread over the surface of food, and Trichosporon has been reported to be present in the maiqu for yellow rice wine. From the statistical analysis results, the main prokaryotic microorganism in daqu was Firmicutes, followed by Actinobacteria, Proteobacteria, Bacteroidetes, Euryarchaeota and Planctomycetaceae. Consistent with literature reports, Firmicutes was the primary prokaryotic microorganism in daqu, and most of these microorganisms are heat resistant, such as Thermoactinomycetes, Bacillus, and Lactobacillus.
Microbial Structure of Daqu of Different Types of Baijiu (15-21)
The Structure of the Eukaryotic Microbes in Daqu
From the statistical analysis of the number of OTUs and the abundance values of eukaryotic microbes for these six kinds of daqu (Figure 2), the results showed that the number of OTUs and abundance values of eukaryotic microbe in daqu of different types of baijiu were different, and the rank order of the number of OTUs was ND˃˃ZD˃QD˃FD˃CD˃JD, and the rank order of the abundance values was QD˃ND˃JD˃ZD˃FD˃CD. The number of OTUs and abundance values of daqu of Nong-xiang baijiu were high, especially 90 OTUs, which was higher than the others. The abundance value of the daqu of Qing-xiang baijiu was 105. The structure of eukaryotic microorganisms in the different types of daqu is shown in Figure 3. The rank order of the relative abundance of eukaryotic microorganisms in Nong-xiang daqu was Aspergillus, Saccharomyces, Eurotium, and Thermomyces, and the relative content of Aspergillus was more than 50%. More than 90% of the total was Saccharomyces in Jiang-xiang daqu. However, differences in the relative abundance of eukaryotic microorganisms were not obvious in Qing-xiang daqu, and the maximum was Xeromyces, which was lower than 20% (Table 2). The eukaryotic microorganisms with the highest relative abundance in Zhimaxiang daqu were Wickerhamomyces and Candida. The eukaryotic microorganisms with the highest relative abundance in Fengxiang daqu were Eurotium and Wickerhamomyces. Overall, the eukaryotic microorganisms with the highest relative abundance in these six types of daqu were Aspergillus, Saccharomyces and some Thermomyces.
Figure 3: Component analysis of relative abundance of eukaryotic microorganisms in the six types of daqu

The Structure of The Prokaryotic Microorganisms in Daqu
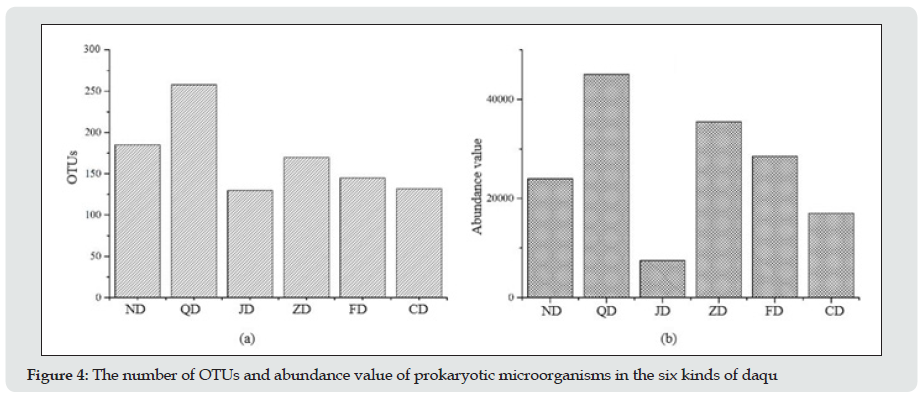
From the statistical analysis of the number of OTUs and abundance values of prokaryotic microorganisms for these six kinds of daqu (Figure 4), the results showed that the rank order of the number of OTUs and abundance values of prokaryotic microorganisms in the daqu of different types of baijiu was QD˃ND˃ZD˃FD˃CD˃JD for the number of OTUs and QD˃ND˃ZD˃FD˃CD˃JD for the abundance values. The number of OTUs and abundance value of the daqu of qing-xiang baijiu were the highest, with 250 OTUs and an abundance value of 4500. Those of the daqu of jiang-xiang baijiu were the lowest.
Figure 4: The number of OTUs and abundance value of prokaryotic microorganisms in the six kinds of daqu
The structure of prokaryotic microorganisms in different types of daqu is shown Figure 5. The prokaryotic microorganisms with the highest relative abundance in Qing-xiang daqu were Staphylococcus, Planifilum, Pediococcus, Lactobacillus and Citrobacter, and the relative contents of Staphylococcus and Planifilum were 55.8% and 22.58%, respectively. The prokaryotic microorganisms with the highest relative abundance in Nongxiang daqu were Weissella, Rheinheimera, Lactobacillus, Bacillus and Pediococcus, and the relative content of Weissella was 38.9%. The prokaryotic microorganisms with the highest relative abundance in Jiang-xiang daqu were Pediococcus, Lactobacillus and Weissella, and these three accounted for more than 90% of the total. The prokaryotic microorganisms with the highest relative abundance in Chi-xiang daqu were Lactobacillus, Citrobacter, Rheinheimera and Lactobacillus, and the relative content of Lactobacillus was more than 70%. The prokaryotic microorganisms with the highest relative abundance in Zhima-xiang daqu were Lentibacillus, Thermoactinomyces, Thermonude, Lactobacillus and Lactobacillus. The prokaryotic microorganisms with the highest relative abundance in Feng-xiang daqu were Lactobacillus, Thermoactinomyces, Thermonude, Lactobacillus and Lactobacillus, and the relative content of Lactobacillus was 65.3%. As shown in Figure 5, the prokaryotic microorganisms with the highest relative abundances in Chinese baijiu daqu were Bacillus, Trichococcus, Pediococcus, Lactobacillus, Weissella, Thermoactinomyces and Thermonude. However, the others, such as Lactococcus, Propionibacterium, and polysporous Actinomycetes, showed great variation in these samples.
Figure 5: Component analysis of the relative abundance of prokaryotic microorganisms in the six types of daqu.

Conclusions
This paper analysed the microbial structure in six different types of Chinese baijiu daqu using NGS.
a) The prokaryotic microorganisms accounted for 73% of the total kinds of microbes, but they accounted for only 38% of the total number. The main eukaryotic microbe in daqu was Ascomycota, followed by Basidiomycota; therein, Ascomycota are mainly moulds and yeasts, and they play a major role in baijiu fermentation. The main prokaryotic microorganism in daqu was Firmicutes, followed by Actinobacteria, Proteobacteria, Bacteroidetes, Euryarchaeota and Planctomycetaceae.
b) The eukaryotic microbes and prokaryotic microorganisms with the highest relative abundances in these six types of Chinese baijiu daqu are shown in Table 3.
References
- Lowes KF, Shearman CA, Payne J, MacKenzie D, Archer DB, et al. (2000) Prevention of yeast spoilage in feed and food by the yeast mycocin HMK. Appl Environ Microbiol 66(3): 1066-1076.
- Kregiel D (2015) Health Safety of Soft Drinks: Contents, Containers, and Microorganisms. Biomed Res Int, p. 1-15.
- Juvonen R, Virkajärvi V, Priha O, Laitila A (2011) Microbiological Spoilage and Safety Risks in Non-Beer Beverages Produced in a Brewery Environment. VTT Tiedotteita-Research, Espoo, Finland.
- Ayala-Zavala JF, del Toro-Sánchez L, Alvarez-Parrilla E, Soto-Valdez H, Martín-Belloso O, et al. (2008) Natural antimicrobial agents incorporated in active packaging to preserve the quality of fresh fruits and vegetables. Stewart Postharvest Review 4: 1-9.
- Cowan MM (1999) Plant products as antimicrobial agents. Clin Microbiol Rev 12: 564-582.
- Kahkha MRR, Amanloo S, Kaykhaii M (2014) Antiaflatoxigenic activity of Carum copticum essential oil. Environ Chem Lett 12(1): 231-234.
- Panizzi L, Flamini G, Cioni PL, Morelli I (1993) Composition and antimicrobial properties of essential oils of four Mediterranean Lamiaceae. J Ethnopharmacol 39(3): 167-170.
- Hili P, Evans CS, Veness RG (1997) Antimicrobial action of essential oils: the effect of dimethylsulphoxide on the activity of cinnamon oil. Lett Appl Microbiol 24(4): 269-275.
- Valsaraj R, Pushpangadan P, Smitt UW, Adsersen A, Nyman U (1997) Antimicrobial screening of selected medicinal plants from India. J Ethnopharmacol 58(2): 75-83.
- Hammer KA, Carson CF, Riley TV (1999) Antimicrobial activity of essential oils and other plant extracts. J Appl Microbiol 86(6): 985-990.
- Binns SE, Purgina B, Bergeron C, Smith ML, Ball L, Baum BR, Arnason JT (2000) Light-mediated antifungal activity of Echinacea extracts. Planta Med 66(3): 241-244.
- Hassawi D, Kharma A (2006) Antimicrobial activity of some medicinal plants against Candida albicans. J Biol Sci 6: 109-114.
- Nzeako BC, Al-Kharousi ZS, Al-Mahrooqui Z (2006) Antimicrobial activities of clove and thyme extracts. Sultan Qaboos Univ Med J 6(1): 33-39.
- López P, Sanchez C, Batlle R, Nerín C (2007) Vapor-phase activities of cinnamon, thyme, and oregano essential oils and key constituents against foodborne microorganisms. J Agric Food Chem 55(11): 4348-4356.
- Pozzatti P, Scheid LA, Spader TB, Atayde ML, Santurio JM, et al. (2008) In vitro activity of essential oils extracted from plants used as spices against fluconazole-resistant and fluconazole-susceptible Candida spp. Can J Microbiol 54(11): 950-956.
- Badiee P, Nasirzadeh A R, Motaffaf M (2012) Comparison of Salvia officinalis L. essential oil and antifungal agents against Candida species. journal of Pharmaceutical Technology and Drug Research 1: 7.
- Bírošová L, Olejníková P, Vaverková Š (2012) Antimicrobial and antimutagenic activities of extracts from different organs of Echinacea angustifolia DC (Asteraceae). J Food Nutr Res 51(4): 201-206.
- Sienkiewicz M, Łysakowska M, Denys P, Kowalczyk E (2012) The antimicrobial activity of thyme essential oil against multidrug resistant clinical bacterial strains. Microb Drug Resist 18(2): 137-148.
- Wendakoon C, Calderon P, Gagnon D (2012) Evaluation of Selected Medicinal Plants Extracted in Different Ethanol Concentrations for Antibacterial Activity against Human Pathogens. Journal of Medicinally Active Plants 1(2): 60-68.
- Abu-Darwish MS, Cabral C, Ferreira IV, Gonçalves MJ, Cavaleiro C, Cruz MT, Al-bdour TH, Salgueiro L (2013) Essential oil of common sage (Salvia officinalis L.) from Jordan: assessment of safety in mammalian cells and its antifungal and anti-inflammatory potential. Biomed Res Int, pp. 538940.
- Giordani R, Regli P, Kaloustian J, Mikaïl C, Abou LH (2004) Portugal, Antifungal Effect of Various Essential Oils against Candida albicans. Potentiation of Antifungal Action of Amphotericin B by Essential Oil from Thymus vulgaris. Phytother. Res 18: 990-995.
- Klaric MS, Kosalec I, Mastelic J, Pieckova E, Pepeljnak S (2007) Antifungal activity of thyme (Thymus vulgaris L.) essential oil and thymol against moulds from damp dwellings. Lett. +Appl. Microbiol 44: 36-42.
- Miladinović D, Miladinović Lj (2000) Antimicrobial activity of essential oil of sage from serbia. Facta Universitatis, Series: Physics, Chemistry and Technology 2(2): 97-100.
- Raal A, Orav A, Arak E (2007) Composition of the essential oil of Salvia officinalis L. from various European countries. Nat Prod Res 21(5): 406- 411.
- Hayouni el A, Chraief I, Abedrabba M, Bouix M, Leveau JY, Mohammed H, Hamdi M (2008) Tunisian Salvia officinalis L. and Schinus molle L. essential oils: their chemical compositions and their preservative effects against Salmonella inoculated in minced beef meat. Int J Food Microbiol 125(3): 242-251.
- Hamidpour M, Hamidpour R, Hamidpour S, Shahlari M (2014) Chemistry, Pharmacology, and Medicinal Property of Sage (Salvia) to Prevent and Cure Illnesses such as Obesity, Diabetes, Depression, Dementia, Lupus, Autism, Heart Disease, and Cancer. J Tradit Complement Med 4(2): 82- 88.
- Khalil R, Li ZG (2011) Antimicrobial activity of essential oil of Salvia officinalis L. collected in Syria. Afr J Biotechnol 10(42): 8397-8402.
- Sookto T, Srithavaj T, Thaweboon S, Thaweboon B, Shrestha B (2013) In vitro effects of Salvia officinalis L. essential oil on Candida albicans. Asian Pac J Trop Biomed 3(5): 376-380.
- Pinto E, Salgueiro LR, Cavaleiro C, Palmeira A, Gonçalves MJ (2007) In vitro susceptibility of some species of yeasts and filamentous fungi to essential oils of Salvia officinalis. Industrial Crops and Products 26: 135- 141.
- Sacchetti G, Maietti S, Muzzoli M, Scaglianti M, Manfredini S, Radice M, Bruni R (2005) Comparative evaluation of 11 essential oils of different origin as functional antioxidants, antiradicals and antimicrobials in foods, Food Chemistry 91: 621-632.
- Jiang Y, Wu N, Fu YJ, Wang W, Luo M, et al. (2011) Chemical composition and antimicrobial activity of the essential oil of Rosemary. Environmental Toxicology and Pharmacology 32: 63-68.
- Moghtader M, Afzali D (2009) Study of the Antimicrobial Properties of the Essential Oil of Rosemary. American-Eurasian J Agric Environ Sci 5(3): 393-397.
- Soković M, Van Griensven LJLD (2006) Antimicrobial activity of essential oils and their components against the three major pathogens of the cultivated button mushroom Agaricus bisporus. Eur J Plant Pathol 116: 211-224.
- Zuzarte M, Gonçalves MJ, Cavaleiro C, Dinis AM, Canhoto JM, et al. (2009) Chemical composition and antifungal activity of the essential oils of Lavandula pedunculata (Miller). Cav Chem Biodivers 6: 1283-1292.
- Koutsoudaki C, Krsek M, Rodger A (2005) Chemical composition and antibacterial activity of the essential oil and the gum of Pistacia lentiscus var. chia. J Agric Food Chem 53: 7681-7685.
- Mazzanti G, Batinelli L, Salvatore G (1998) Antimicrobial properties of the linalool-rich essential oil of Hyssopus officinalis var. decumbens (Lamiaceae). Flavour Frag J 13: 289-294.
- Merali S, Binns S, Paulin-Levasseur M, Ficker C, et al. (2003) Antifungal and antiinflammatory activity of the genus Echinacea. Pharmaceutical Biology 41(6): 412-420.
- Kumar KM, Ramaiah S (2011) Pharmacological importance of Echinacea purpurea. International Journal of Pharma and Bio Sciences 2(4): 304- 314.
- Singh N (2010) A comparison of both water and ethanol extracts prepared from Echinacea purpurea and Echinacea angustifolia on the response to Influenza A/PR/8/34 infection in mice. Thesis master of science, Iowa State University.
- Stoll A, Renz J, Brack A (1950) Isolierung und Konstitution des echinacosides, eines glykosids aus den wurzeln von Echinacea angustifolia D.C. Helv Chim Acta 33: 1877-1893.
- Jin M, Huang Q, Zhao K, Shang P (2013) Biological activities and potential health benefit effects of polysaccharides isolated from Lycium barbarum L. Int J Biol Macromol 54:16-23.
- Schepetkin IA, Quinn MT (2006) Botanical polysaccharides: macrophage immunomodulation and therapeutic potential. Int Immunopharmacol 6(3): 317-333.
- Sun Y (2011) Structure and biological activities of the polysaccharides from the leaves, roots and fruits of Panax ginseng CA Meyer: An overview. Carbohyd Polym 85(3): 490-499.
- Asressu KH, Tesema TK (2014) Chemical and antimicrobial investigations on essential oil of Rosmarinus officinalis leaves grown in ethiopia and comparison with other countries. J App Pharm 6(2): 132-142.
- Steinmuller C, Roesler J, Grottrup E, Franke G, Wagner H, et al. (1993) Polysaccharides Isolated From Plant Cell Cultures Of Echinacea Purpurea Enhance The Resistance Of Immunosuppressed Mice Against Systemic Infections With Candida albicans and Listeria Monocytogenes. International Journal of Immunopharmacology 15: 605-614.

Top Editors
-

Mark E Smith
Bio chemistry
University of Texas Medical Branch, USA -

Lawrence A Presley
Department of Criminal Justice
Liberty University, USA -

Thomas W Miller
Department of Psychiatry
University of Kentucky, USA -

Gjumrakch Aliev
Department of Medicine
Gally International Biomedical Research & Consulting LLC, USA -

Christopher Bryant
Department of Urbanisation and Agricultural
Montreal university, USA -

Robert William Frare
Oral & Maxillofacial Pathology
New York University, USA -

Rudolph Modesto Navari
Gastroenterology and Hepatology
University of Alabama, UK -

Andrew Hague
Department of Medicine
Universities of Bradford, UK -

George Gregory Buttigieg
Maltese College of Obstetrics and Gynaecology, Europe -

Chen-Hsiung Yeh
Oncology
Circulogene Theranostics, England -
.png)
Emilio Bucio-Carrillo
Radiation Chemistry
National University of Mexico, USA -
.jpg)
Casey J Grenier
Analytical Chemistry
Wentworth Institute of Technology, USA -
Hany Atalah
Minimally Invasive Surgery
Mercer University school of Medicine, USA -

Abu-Hussein Muhamad
Pediatric Dentistry
University of Athens , Greece

The annual scholar awards from Lupine Publishers honor a selected number Read More...