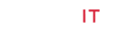Lupine Publishers Group
Lupine Publishers
Menu
ISSN: 2637-4676
Research Article(ISSN: 2637-4676) 
Allele Mining for the Reported Genes Governing the Yield Related Traits in a Set of Rice Germplasm Using PCR-Based Markers Volume 3 - Issue 4
Bidanchi T, M Sangma, VK Khanna*and W Tyagi
- School of Crop Improvement, Central Agricultural University, India
Received: June 22, 2018; Published: July 12, 2018
Corresponding author: VK Khanna, School of Crop Improvement, Central Agricultural University, Barapani, Umiam-793103, Meghalaya, India
DOI: 10.32474/CIACR.2018.03.000169
Abstract
The present study was an initial attempt towards enhancing grain yield in rice through molecular breeding approach by allele mining. Forty one diverse rice genotypes were used for allele mining study with an objective to detect among them superior alleles for rice grain dimensions viz. grain length, grain width, grain thickness, grain size and grain weight. SSR markers associated with reported genes for these traits were used for PCR amplification. It was found that SSR primer RDD 1-2 can be used for identifying alleles enhancing grain length from diverse genomic sources for crop improvement. The allele A (600 bp) from Rdd 1 can be used in breeding programmes to enhance grain length in rice. The genotypes viz. Hans raj, Pant dhan-18, Pant dhan-10, Narendra, Pant dhan-12, Pant dhan-4, ARR-09, Shahsarang, IVT-ASG-2712, IVT-ASG-2701, AVT-I-ASG-2602, AVT-I-ASG-2609 and IVT-ASG-2705 identified carrying this allele can be used as a source for large grain length in future. The primers RM 478 for grain thickness and RM 574-2 for grain width can be used in marker assisted breeding programmes. Allele A (205 bp) of RM 478 and allele B (240 bp) of RM 574-2 associated with low grain weight and grain width, respectively can be used for rejection at seedling stage in marker assisted breeding programme. This study also identified novel alleles (alleles distinct from those reported in biparental programme) which could be significant in a larger panel of rice genotypes.
Keywords: Allele mining; rice;Oryza sativa; O. nivara; SSR primers
Introduction
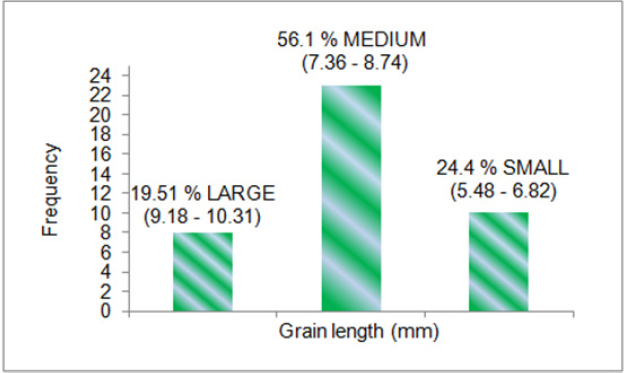
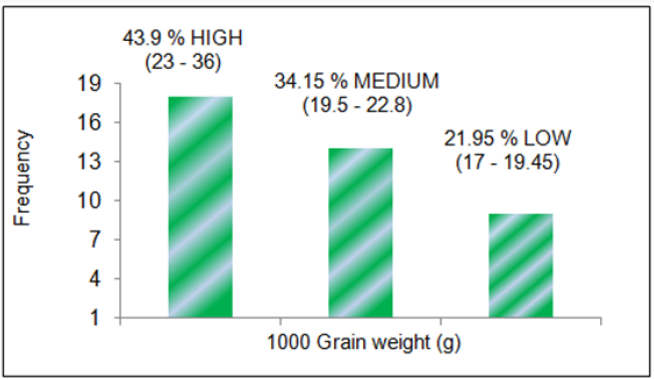
A panel of 41 rice genotypes consisting of the released cultivars of O. sativa, landraces and nineteen accessions from AVT, AVIT and IVT series and two accessions RWR-19 and RWR-125 from wild species O. nivara were used for allele mining study for grain traits contributing to grain yield. The mean data for grain length, grain width and 1000 grain weight was taken for all the genotypes used for allele mining study. The 41 rice genotypes were categorized into three groups in order of their grain dimensions and 1000 grain weight. The genotypes consisted of 39.02 % large (2.49-3.62), 51.22 % medium (2.01-2.44) and 9.76 % small (1.85-1.97) for grain width in mm; 19.51 % large (9.18-10.31), 56.1 % medium (7.36-8.74) and 24.4 % small (5.48-6.82) for grain length in mm; 43.9 % high (23- 36), 34.15 % medium (19.5-22.8) and 21.95 % small (17-19.45) for 1000 grain weight in grams. F-test was conducted for analysis of variance in phenotypic traits between the genotypes. Significant variation at 5% level was observed for grain length (F=9.61) and grain width (F=6.10). Allele mining for grain yield contributing traits viz. grain length, grain weight and grain width was done for the reported gene Rdd1 and the QTLs tightly associated with the traits with an aim to identify alleles of genes conferring the grain yield traits. The genes/QTLs associated with these three-grain yield contributing traits viz. grain length, grain width and 1000 grain weight targeted in this study included Rdd 1 (Rice dof daily fluctuations 1), gw8 (Grain width 8), gt7 (Grain thickness 7), gw 8.1 (grain weight 8.1), G S5 (Grain size 5), and gw7 (Grain weight 7), all previously reported to be associated with grain yield traits in biparental studies (except Rdd 1).
Scoring of PCR products was done through gel electrophoresis by comparing the size of the bands of the PCR products using a gel documentation system. The markers showing bands of similar positions in gels for all the genotypes were monomorphic markers while those markers showing bands at different position were polymorphic markers. All the genotypes that show products of similar size with that of expected amplicon size for the reported trait will carry the desired allele for grain yield and likewise the alleles for all the target regions were identified. PCR-based allele mining for these traits was performed in a panel of 41 genotypes using 10 microsatellite markers reported to be tagged with these genes/QTLs. Ten microsatellite markers were used for allele mining; out of which nine markers (RDD1-2, RM 234, RM 478, RM 23201, RM 502, RM 574-1, RM 574-2, RM 593-1 and RM 593-2) amplified. Out of these nine markers, RM 23201 was monomorphic and the rest were polymorphic. Maximum number of four alleles was found for marker RM 478 and RM 574-2, three alleles for RDD I-2 and RM 234 and two alleles for RM 502, RM 574-1, RM 593-1 and RM 593-2 when scored in agarose gel. Chi square test was conducted to study the association of the markers with grain traits [1]. Each of the alleles observed in the top 15 and bottom 15 genotypes were tested for the consistency of their distribution with the whole population. Significant associations were observed for allele A in 15 largest genotypes (Χ2=3.94, p=0.05) and B allele (Χ2=4.51, p=0.03) in the 15 smallest length genotypes for primer RDD 1-2. Allele A was found to be associated with large grain length and allele B was found to be associated with small grain length. The genotypes viz. Hans raj, Pant dhan-18, Pant dhan-10, Narendra, Pant dhan-12, Pant dhan-4, ARR-09, Shahsarang, IVT-ASG-2712, IVT-ASG-2701, AVT-IASG-2602, AVT-IASG-2609 and IVT-ASG-2705 were found to contain A allele found to be associated with large grain length for RDD 1-2 marker. Significant association was observed in marker RM 478 with grain weight and for distribution of A allele in the bottom 15 genotypes (lowest grain weight) with the distribution in the overall population (Χ2=5.08, p=0.02) and for primer RM 574-2, distribution of B allele (Χ2=4.3, p=0.08) in the bottom 15 genotypes (smallest grain width) was observed to be significant.
From this present experiment, hybrid plants from inter-specific crosses can be used for backcrossing in future plant breeding programmes. Crosses can also be attempted for genotypes that failed during this experiment. Primer RDD 1-2 can be used for identifying alleles enhancing grain length from diverse genomic sources for crop improvement [2]. The allele A (600 bp) from Rdd 1 can be used in breeding programmes to enhance grain length in rice. The genotypes viz. Hans raj, Pant dhan-18, Pant dhan-10, Narendra, Pant dhan-12, Pant dhan-4, ARR-09, Shahsarang, IVTASG- 2712, IVT-ASG-2701, AVT-IASG-2602, AVT-IASG-2609 and IVT-ASG-2705 were identified to be carrying this allele can be used as a source for large grain length in future. The primers RM 478 for grain thickness and RM 574-2 for grain width can be used in marker assisted breeding programmes. Allele A (205 bp) of RM 478 and allele B (240bp) of RM 574-2 associated with low grain weight and grain width, respectively can be used for rejection at seedling stage in marker assisted breeding programme. This study also identified novel alleles (alleles distinct from those reported in biparental programme), which could be significant in a larger panel of rice [3].
Materials and Methods
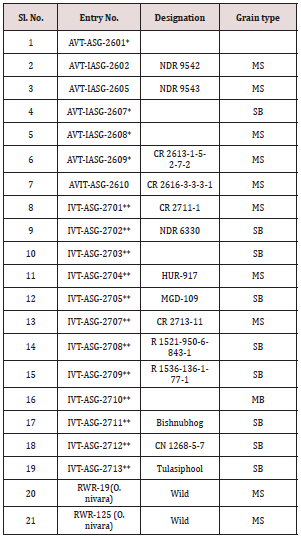
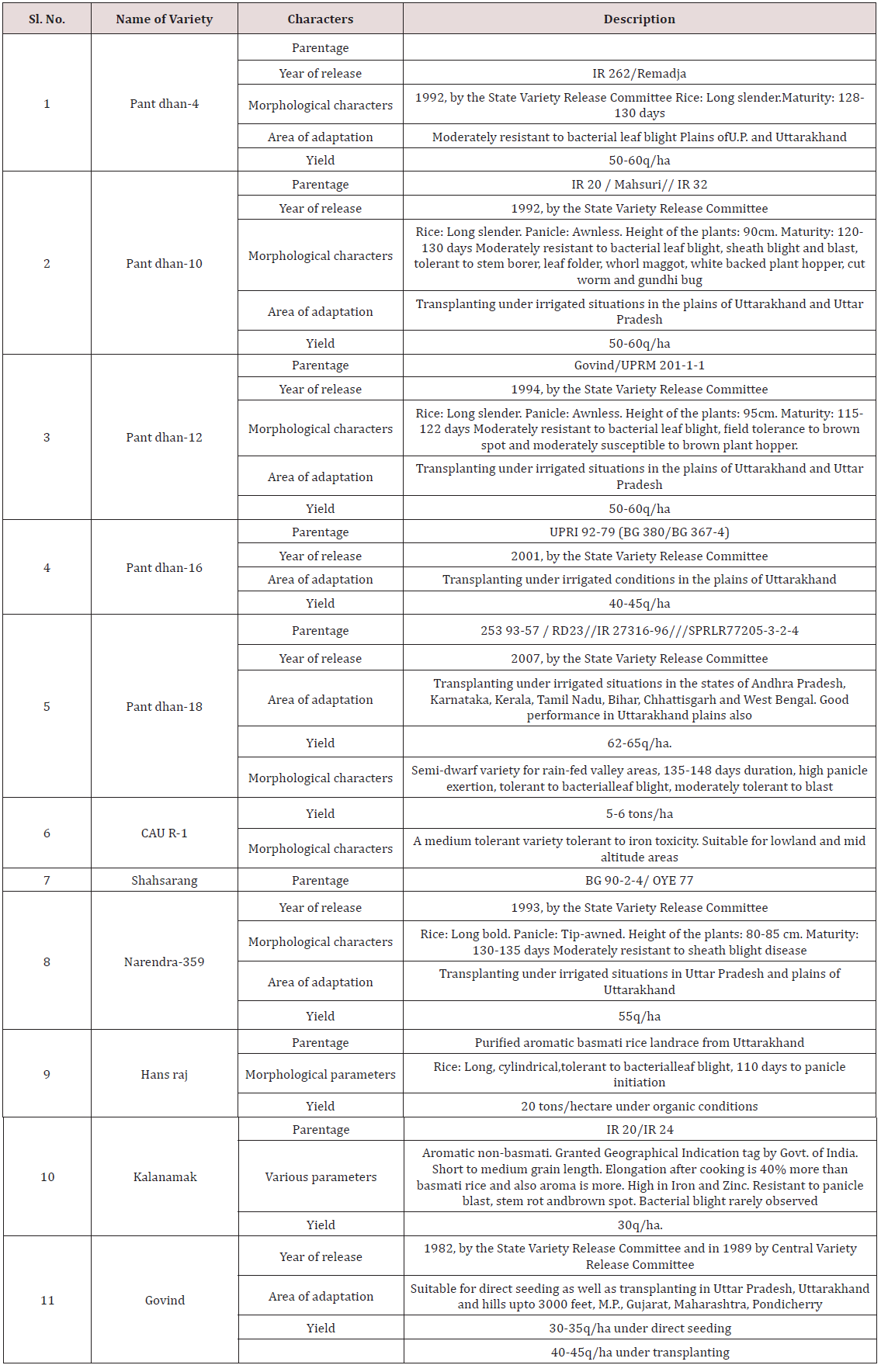
A panel of different rice genotypes including released cultivars of O. sativa i.e., IR-1552 (LR-55), Govind, CAU R-1, Kalanamak, Hansraj, Pant dhan-4, Pant dhan-10, Pant dhan-12, Pant dhan-16, Pant dhan-18, Narendra-359, landraces like ARR-09, LR-1 (Balwai), LR-5 (Laljagli), LR-23 (Sabhagidhan), LR-71 (Basmati paddy), LR- 77 (BR-1),Shahsarang, Bhasphool (LR-2), Chakaoporieton (LR- 26), and nineteen accessions from AVT and IVT series and two accessions RWR-19 and RWR-125 from wild species O. nivara were used for allele mining. The experimental material seed of Pant dhan was obtained from Pantnagar and seed of AVT, AVIT and IVT series have been procured from DRR (Directorate of Rice Research), Hyderabad (Tables 1-3) [4].
*- Advanced varietal trial; **- Initial varietal trial; SB: Short bold; MS: Medium slender/Mild scented; LS: Long slender; SS: Strong scented.
Statistical analysis of seed parameters for grain size variance in 41 rice genotypes
The grain size of rice is contributed collectively by a complex of yield related traits of which grain length, grain width and grain weight also play a major role. Given prior concern to these three traits, a set of data for average grain length, grain width and grain weight has been taken for all the genotypes used for allele mining.
Phenotypic analysis/Analysis of variance
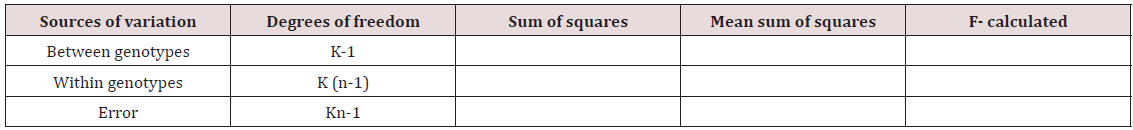
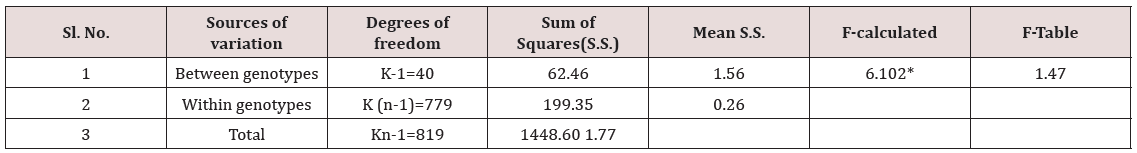
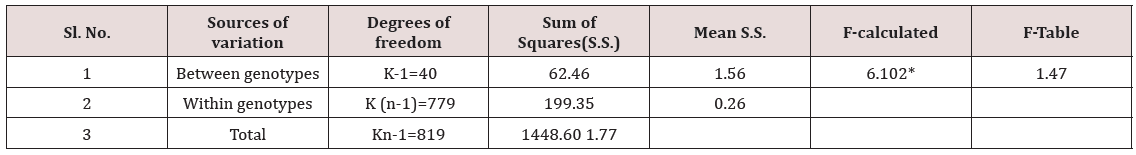
The frequency distribution in percent for grain length, grain width and grain weight was determined for large, medium and small grains in the population of 41 genotypes. Analysis of variance for significance of phenotypic variations for grain length and grain width of 41 genotypes was studied through F-test (Table 4).
Germination and sowing of seeds
Sowing of seeds used for allele mining study was done in the month of February 2014 (Table 4). Prior to sowing, seeds of all the genotypes were first allowed to germinate in sterilized Petri plates containing filter papers moistened with water. Upon germination, the seeds were transferred to shallow plastic trays and the genotypes were grown in rows with each genotype in a single row having at least twenty seedlings per row [5]. The soil was finely prepared free of clods and was mixed with farm yard manure. The trays containing the plants were kept inside the greenhouse and the plants were maintained by regular watering and also weeding as and when required.
Molecular Characterization
SSR analysis for polymorphism
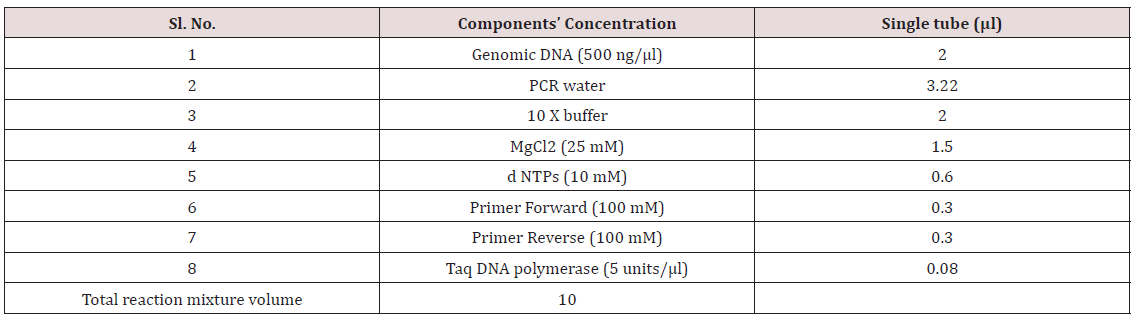
Rice dof daily fluctuations1 (Rdd1), Grain width5 (GS5), Grain thickness (gt7) and Grain weight8 (GW8) previously reported to be linked to the grain traits/QTLs were used for allele mining studies in different rice genotypes. Forty one rice genotypes including two accessions of O. nivara were used for allele mining studies to detect the existing alleles among the genotypes for the given traits. Markers for these reported genes were obtained from Gramene marker database. Genomic DNA from rice leaves of all the genotypes were isolated using CTAB method of Doyle [3]. The DNA samples were diluted to obtain a working standard according to the concentration and the quality of the DNA obtained. The PCR reaction conditions for primers were standardized to obtain good quality bands and were used as the standard reaction conditions throughout the whole PCR amplification process [6].
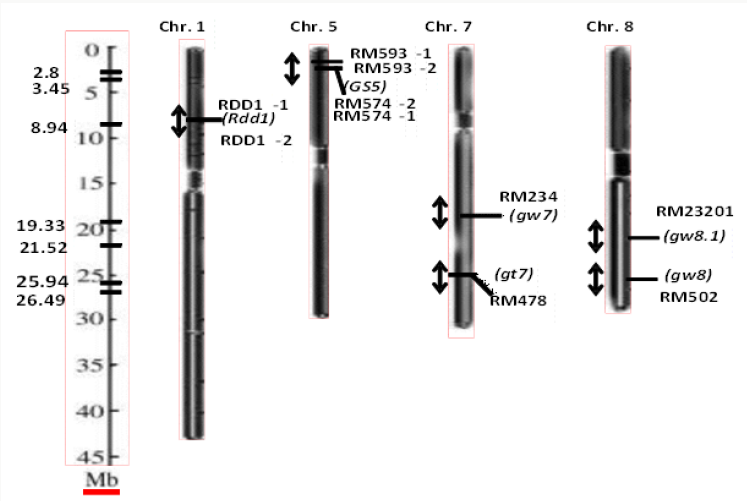
The three yield contributing traits namely grain length, grain width and grain weight was targeted in this study. The gene Rdd1 has been reported to be indirectly related to increase in grain size in rice. Since increase in grain size is constitutively contributed by grain traits which include grain length, the RDD1 markers associated with this gene has been used in allele mining for grain length trait. Gs 5 is a reported QTL which has been known to associate positively with grain width and thus contributing to grain size and the RM markers reported to be associated with this gene and flanking at the regions near it were used for allele mining study. Another QTL, gw 8, known to positively control grain width and QTLs; gt7, gw 8.1 and gw 7 for grain weight were targeted for allele mining in the present study (Figure 1).
Figure 1: Chromosome map showing genes/QTLs (italicized) associated with grain yield traits and the position of the markers (designated as RDD 1 and RM) linked to these traits respectively, in rice. The scale of the maps in megabase is shown on the left axis.
Isolation of Genomic Dna from Plant Tissue
Genomic DNA was isolated from genotypes used for allele mining. Tender leaves from young rice seedlings were used for the extraction of genomic DNA. Genomic DNA from rice leaves was isolated using CTAB method of Doyle [3]. Young actively growing leaves of 15 days-old plants were collected and used for DNA extraction.
Procedure for DNA extraction
About 2g of young fresh plant leaf tissue was rapidly frozen in liquid nitrogen and ground to a fine powder in a pre-chilled sterile pestle and mortar. The frozen powder was transferred to a 2ml tube. 1ml of CTAB extraction buffer was added to each tube and tubes were inverted several times with gentle shaking. The tubes were incubated at 65 0C in a water bath for 1 hr. Contents were mixed by inverting the tubes after every 10-15 minutes. The tubes were cooled to room temperature and 1 ml of chloroform: isoamyl alcohol in the ratio of 24:1 was added to each. The contents were mixed by inverting the tubes several times. The tubes were centrifuged at 10,000 rpm for 10 minutes at 4 0C. The upper clear aqueous supernatant was removed and transferred to fresh tubes. 1ml of pre-chilled isopropanol was added to each tube and the contents were mixed by inverting the tubes several times and the tubes were kept overnight at -20 0C. Although DNA precipitation began as soon as isopropanol was added, it was kept overnight for better and complete precipitation. The samples were centrifuged for 10 minutes at 5,000 rpm at 4 0C. The solution was poured without disturbing the DNA pellet at the bottom of the tubes. 1.5ml of 70% ethanol was added and then centrifuged at 5,000rpm for 5 minutes at 4 0C. Supernatant was decanted. Pellet was air dried for 15-20 minutes. 50μl of TE buffer was added to the pellet [7].
DNA quantification
The quantification of DNA was done by staining DNA with ethidium bromide after electrophoresis in 0.8% agarose gel at 100V for 1 hour in TBE buffer (0.04M Tris borate, 0.001 M EDTA, pH 8.0) using known DNA concentration standard.
Dilution of DNA samples
A part of the DNA sample of each genotype was taken and dilution of each sample was made with appropriate amount of TE buffer to yield a working concentration and stored at -20°C for further use in PCR amplification.
Polymerase chain reaction (PCR)
Dilution of SSR markers: The SSR markers which were in lyophilized state were centrifuged and diluted with TE buffer accordingly to yield 100pmol/μl stock concentration and from the stock, working concentration of 10pmol/μl was made by diluting at 1:9 ratio of stock with TE buffer. The primer stock was immediately transferred to -20 °C to prevent degradation during longer duration storage and the working standard was kept in 4 °C for use in the study.
Standardization of PCR conditions
Standardization of annealing temperature: Annealing temperatures for each of the primers was standardized by PCR reaction at least three different PCR conditions. The reaction condition giving amplification with good bands were taken as the standard for all the following experiments.
PCR conditions for markers used for allele mining
Primary denaturation: 95 °C for 3 minutes
Denaturation: 95 °C for 0.30 minutes
Annealing: 55 °C for 0.45 minutes
Extension: 72 °C for 0.40 minutes
Final extension: 72 °C for 2 minutes
Number of cycle: 32
Allele mining of reported genes/ QTL for yield related grain traits in rice using SSR markers
In this study, 41 diverse rice genotypes were used for allele mining study with an objective to detect among them superior alleles for rice grain dimensions viz. grain length, grain width, grain thickness, grain size and grain weight. SSR markers associated with reported genes for these traits were used for PCR amplification Micro satellite markers linked to reported genes/QTL for grain yield traits were used to detect polymorphism and to detect the yield enhancing alleles among 41 rice genotypes. PCR amplification was done with a set of 10 primers (Tables 5 & 6) and analysis for polymorphism was done through electrophoresis in 3% agarose gel. Scoring of PCR products was done through gel electrophoresis by comparing the size of the bands of the PCR products using a gel documentation system. The markers showing bands of similar positions in gels for all the genotypes were monomorphic markers while those markers showing bands at different position were polymorphic markers. All the genotypes that show products of similar size with that of expected amplicon size for the reported trait will carry the desired allele for grain yield and likewise the alleles for the entire target regions were identified [8].
Association between phenotype and genotype
The average grain length, grain width and 1000 grain weight distribution in the genotypes carrying different alleles was determined by checking whether the error bars referring to confidence interval showed overlapping or not. The 41 genotypes were further categorized into top and bottom groups of 15 genotypes based on grain length, grain width and grain weight. Chi square analysis was performed to study the association between markers with grain traits and allelic association with the trait for each of the sub-groups.
Results and Discussion
Phenotypic analysis for yield related traits
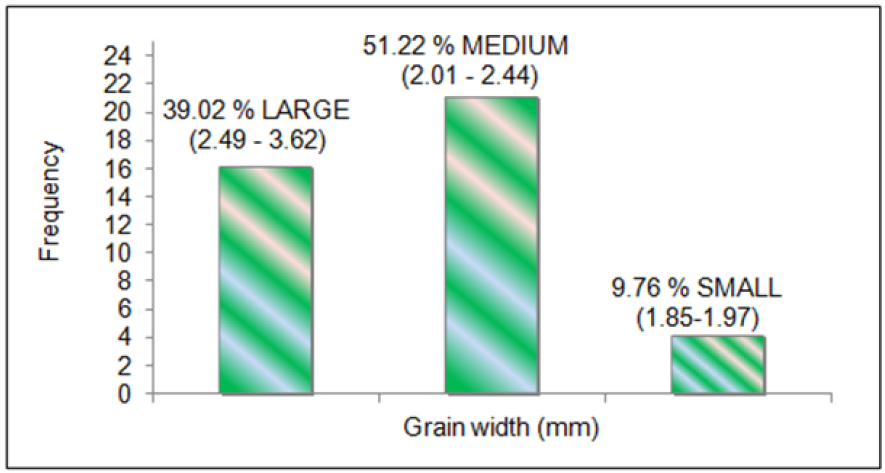
Phenotypic variation was observed for grain dimensions in all the genotypes. The mean data for grain width (Figure 2), grain length (Figure 3) and 1000 grain weight (Figure 4) was taken for all the 41 genotypes used for allele mining study. The genotypes with larger grain width (mm) were LR-1 (3.62), LR-71 (3.07), Pant dhan- 16 (2.85), IVT-2712 (2.79) and RWR-19 (2.77). Smaller grain width (mm) was observed in Kalanamak (2.01), AVT-2607 (2.0), Hans raj (1.97), AVT-2701 (1.94), IVT-2703 (1.904) and AVT-2609 (1.85). The panel consisted of 39.02 % large (2.49-3.62), 51.22 % medium (2.01-2.44) and 9.76 % small (1.85-1.97) for grain width (in mm) trait. The genotypes with larger grain length (mm) were Hans raj (10.31), Pant dhan-18 (9.83), Pant dhan-10 (9.7), Narendra (9.63), Pant dhan-12 (9.53), RWR-125 (9.27), Pant dhan-4 (9.23) and Govind (9.12). Smaller grain length (mm) was observed for genotypes IVT-ASG-2708 (6.41), Kalanamak (6.35), IVT-ASG-2711 (6.15), IVT-ASG-2607 (5.92), IVT-ASG-2713 (5.71), IVT-ASG-2703 (5.69), LR-71 (5.59) and LR-77 (5.48). The genotype panel consisted of 19.51 % large (9.18-10.31), 56.1 % medium (7.36-8.74) and 24.4 % small (5.48-6.82) for grain length (in mm).The genotypes having the highest grain weight (g) were LR-1 (36.04), Pant dhan- 18 (27.97), Pant dhan-4 (26.39), LR-23 (25.32), RWR-125 (25.05), Narendra (24.2), Pant dhan -12 (23.97) and Pant dhan-16 (23.39). Lower grain weight (g) was observed for genotypes, Kalanamak (12.17), LR-77 (12.14), AVT-IASG-2609 (12.00), IVT-ASG-2713 (11.96), AVT-IASG-2607 (10.74) and IVT-ASG-2703 (10.72). The genotype panel consisted of 43.9 % high (23-36), 34.15 % medium (19.5-22.8) and 21.95 % small (17-19.45) for 1000 grain weight (g). The phenotypic variance observed for the 41 rice genotypes were found to be significant at 5% level for grain width (F=9.61) and grain length (F=6.1).
Allele mining for genes/QTLs
Alleles are the alternative DNA sequences at the same physical locus which may or may not result in different phenotypic traits. Increasing demand of rice varieties with higher yield is making scientists work for the identification of superior and novel alleles to be used in breeding programmes. Allele mining is another very important molecular approach towards identifying and bringing together useful traits from diverse genetic sources and introgression these useful alleles into a common recipient promising cultivar lacking one or two superior alleles for some agronomic traits. Molecular markers are effectively being used to detect the variant alleles present in diverse genotypes or cultivars which are phenotypically different from each other for the traits in question. Microsatellite markers or SSR markers are PCR-based markers that specifically bind to the complementary sequence of the plant genomic DNA during PCR reaction. Through allele mining novel genes could be identified from diverse sources and used in improvement of crops for specific traits. Allele mining for grain yield related traits in large number of genotypes/ cultivars would help in identifying new alleles of the genes known for the target trait and their sources. It can also provide an insight into molecular basis of novel trait variations and identify the nucleotide sequence changes associated with superior alleles [9].
The quantitative trait locus (QTL) mapping has contributed to a better understanding of the genetic basis of many agronomically important traits such as grain yield. Grain yield is a quantitatively inherited trait which is the result collectively contributed by several individual grain traits. Of these, grain length, grain width and grain weight are some of the important yield contributing grain traits in rice. Today, a number of genes/QTLs associated with these traits have been identified, sequenced, mapped and many associated markers to these QTLs have been developed recently. In this study, a set of 41 diverse genotypes including two accessions viz. RWR- 19 and RWR-125 from wild species O. nivara were used for allele mining using nine SSR markers tagged to reported genes/QTLs. Out of the nine primers viz. RDD 1-2, RM 234, RM 478, RM 23201, RM 502, RM 574-1, RM 574-2, RM 593-1 and RM 593-2 used in the present study, RM 23201 was found to be monomorphic and the rest eight primers were polymorphic.
Association between genotype and phenotype
Chi square test was conducted to study the association of the markers with grain traits. Each of the alleles observed in the top 15 and bottom 15 genotypes were tested for the consistency of their distribution with the whole population. Significant association (Χ2=3.94, p=0.05) was observed for allele A in the 15 genotypes having large grain length for primer RDD1-2. Allele Bfor Rdd1 gene too showed significant association (Χ2=4.51, p=0.03). Allele A was found to be associated with large grain length and allele B was found to be associated with small grain length. Significant association was observed for marker RM 478 with grain weight and for the distribution of A allele in the bottom 15 genotypes (lowest grain weight) with the distribution in the overall population (Χ2=5.08, p=0.02) and for primer RM574-2, distribution of B allele (Χ2=4.3, p=0.08) in the bottom 15 genotypes (smallest grain width) was found to be significant.
Rdd 1 (Rice dof daily fluctuations1) Dof gene in rice has been reported to be associated with plant growth, grain size and flowering timein rice. In a study conducted by Iwamoto et al. (2009), transgenic plants carrying full length Rdd 1cDNA driven by constitutive promoter were produced in order to study the role of Rdd1 and in - vivo function of Dof gene in plants. The two types of transgenic plants were produced viz. RDD 1-S, transgenic containing sense strand and RDD 1-AS with anti-sense strand. Differences in the grain length, grain width and grain weight were observed between RDD 1-AS and RDD 1-S plants when subjected under varying photoperiod conditions [4]. When the grain sizes of the two transgenic types of plants were compared with the wild type plants, there was significant decrease in grain length, grain width and decrease in 1000 grain weight in transgenics carrying antisense gene (AS1=4.67 and AS2=4.54). In our study, we used previously reverse RDD-1 primer and designed a new forward primer giving an amplicon size of 620 bp. The three alleles obtained in the present study with sizes of 600, 620 and 600’490 base pairs were designated as A, B and C, respectively. Significant association (Χ2=3.94, p<0.05) was observed between the grain length and the allele A (600bp) for large grain length and allele B (620 bp) associated with small grain length (Χ2=4.51, p<0.05). The allele A for Rdd1 can be used in breeding for large grain length. In our study, however, a set of 41 genotypes having large grain length ranging from 8.27-10.31mm and small grain length of 5.48-7.52 mm were used. This phenotypic variation is distinct from the WT genotype Nipponbare (4.94mm) used by Iwamoto [4]. The allele A though similar in size could be different due to nucleotide variation. In order to rule out that possibility, sequencing of the allele from a panel of genotypes showing large grain length will have to be done.
GS 5 (Grain size 5) QTL has been reported to be a positive regulator of grain width and flanked by markers RM 593 and RM 574. Over-expression of GS5 promotes cell division resulting in an increase of grain width [6]. Other genes/QTLs in rice are Grain weight 7 (gw 7) reported to be associated with 1000 grain weight located at chromosome 7 in population derived from O. sativa and O. grandiglumis. Phenotypic variation of 13.3% was observed in populations having allele for gw 7 from Caipo (Aluko, 2003). Grain width 8 (gw 8) at chromosome 8 has been reported to have negative effect on grain width with decrease in grain width in populations resulting from separate crosses made between HJX 74 x Anmol 3 and HJX 74 x Basmati 370, with possible introgression of the recessive allele from Anmol 3 and Basmati 370 [7,8]. No significant association was observed for primers reported for Grain weight 8.1 (gw 8.1) [9] QTL in our panel of 41 genotypes. Grain thickness 7 (gt 7), a QTL for grain thickness with linked marker RM 478 at chromosome 7¸ has been reported to be associated positively with grain thickness in HG 101 which is a near isogenic line obtained from O. sativa (Hwaseongbyeo) x O. grandiglumis. T [8].
he increased grain length in mm (5.53), grain width in mm (3.08) and 1000 grain weight in grams (26.3) was observed for the NILs as compared to the recipient parent, Hwaseongbyeo, with grain length (5.03 mm), grain width (2.84mm) and 1000 grain weight (21.5g). In the present study, significant association of primer RM 478 was observed and the allele A (205bp) was significantly associated with low 1000 grain weight. The genotypes used in the present study have highest 1000 grain weight ranging from 20.16-36.04 grams and lowest ranging from 10.72-15.59 grams. Since grain yield is quantitative in nature, it is a complex trait and the markers tagged with the genes/QTLs may not show association in a diverse set of genotypes. Also the effect of individual genic components to grain yield may be very low as grain yield is a trait caused by cumulative action of many related components. Our study however, suggests that the allele A (600 bp) from Rdd 1 can be used in breeding programmes to enhance grain length in rice. The genotypes viz. Hans raj, Pant dhan-18, Pant dhan-10, Narendra, Pant dhan-12, Pant dhan-4, ARR-09, Shahsarang, IVT-ASG-2712, IVT-ASG-2701, AVT-IASG-2602, AVT-IASG-2609 and IVT-ASG-2705 have been identified to be carrying this allele. These can be used as a source of large grain length in future. The primers RM 478 for grain thickness and RM 574-2 for grain width can be used in marker assisted breeding programmes. Allele A (205 bp) of RM 478 and allele B (240 bp) of RM 574-2 associated with low grain weight and grain width, respectively can be used for rejection at seedling stage in marker assisted breeding programme (Table 7). This study also identified novel alleles (alleles distinct from those reported in biparental programme); which could be significant in a larger panel of rice genotypes (Figure 2-4).
Discussion
Phenotypic analysis for yield related traits
Phenotypic variation was observed for grain dimensions in all the genotypes. The mean data for grain width (Figure 5), grain length (Figure 6) and 1000 grain weight (Figure 7) was taken for all the 41 genotypes used for allele mining study. The genotypes with larger grain width (mm) were LR-1 (3.62), LR-71 (3.07), Pant dhan- 16 (2.85), IVT-2712 (2.79) and RWR-19 (2.77). Smaller grain width (mm) was observed in Kalanamak (2.01), AVT-2607 (2.0), Hans raj (1.97), AVT-2701 (1.94), IVT-2703 (1.904) and AVT-2609 (1.85). The panel consisted of 39.02 % large (2.49-3.62), 51.22 % medium (2.01-2.44) and 9.76 % small (1.85-1.97) for grain width (in mm) trait. The genotypes with larger grain length (mm) were Hans raj (10.31), Pant dhan-18 (9.83), Pant dhan-10 (9.7), Narendra (9.63), Pant dhan-12 (9.53), RWR-125 (9.27), Pant dhan-4 (9.23) and Govind (9.12). Smaller grain length (mm) was observed for genotypes IVT-ASG-2708 (6.41), Kalanamak (6.35), IVT-ASG-2711 (6.15), IVT-ASG-2607 (5.92), IVT-ASG-2713 (5.71), IVT-ASG-2703 (5.69), LR-71 (5.59) and LR-77 (5.48). The genotype panel consisted of 19.51 % large (9.18-10.31), 56.1 % medium (7.36-8.74) and 24.4 % small (5.48-6.82) for grain length (in mm). The genotypes having the highest grain weight (g) were LR-1 (36.04), Pant dhan- 18 (27.97), Pant dhan-4 (26.39), LR-23 (25.32), RWR-125 (25.05), Narendra (24.2), Pant dhan -12 (23.97) and Pant dhan-16 (23.39). Lower grain weight (g) was observed for genotypes, Kalanamak (12.17), LR-77 (12.14), AVT-IASG-2609 (12.00), IVT-ASG-2713 (11.96), AVT-IASG-2607 (10.74) and IVT-ASG-2703 (10.72) (Tables 8 & 9). The genotype panel consisted of 43.9 % high (23-36), 34.15 % medium (19.5-22.8) and 21.95 % small (17-19.45) for 1000 grain weight (g). The phenotypic variance observed for the 41 rice genotypes were found to be significant at 5% level for grain width (F=9.61) and grain length (F=6.1).
Figure 5: SSR profile of grain length generated by primer RDD1-2 in 41 rice genotypes. The numbers at the top refer to the genomic DNA code as mentioned in Table 4.2. Alleles are indicated at the bottom. Allele A=approximately 600 bp; allele B=620 bp; B’C=620’ 490 bp; X=no amplification. 1kbp ladder is indicated at the right of the gel.
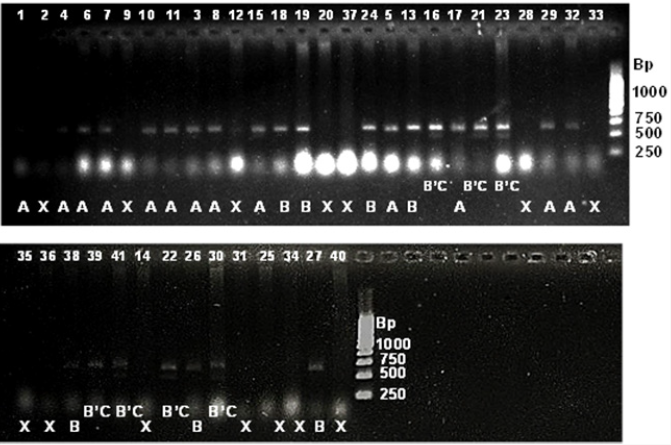
Figure 6: SSR profile of grain weight generated by primer RM 234 in 41 rice genotypes. The numbers at the top refer to the genomic DNA code as mentioned in Table 4.2. Alleles are indicated at the bottom. Allele A=approximately 110 bp; allele B=125 bp; allele C=155; X=no amplification. 100 bpladder is indicated to the right of the gel.
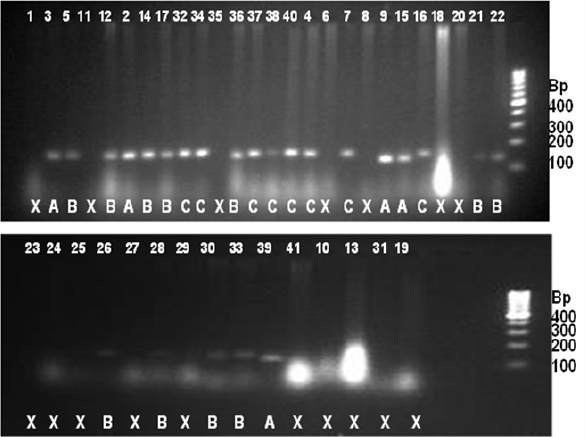

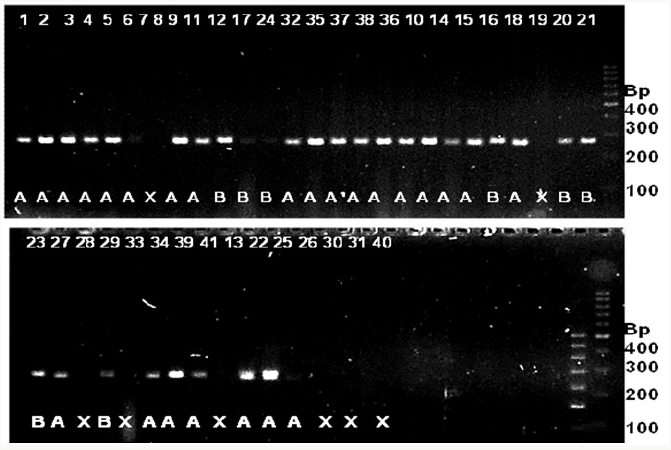
Figure 7: SSR profile of grain width generated by primer RM 502 in 41 rice genotypes. The numbers at the top refer to the genomic DNA code as mentioned in Table 4.2. Alleles are indicated at the bottom. Allele A=approximately 255 bp; B=170 bp; X=no amplification. 100 bpladder is indicated to the right of the gel.
Allele mining for genes/QTLs
Alleles are the alternative DNA sequences at the same physical locus which may or may not result in different phenotypic traits. Increasing demand of rice varieties with higher yield is making scientists work for the identification of superior and novel alleles to be used in breeding programmes. Allele mining is another very important molecular approach towards identifying and bringing together useful traits from diverse genetic sources and introgression these useful alleles into a common recipient promising cultivar lacking one or two superior alleles for some agronomic traits. Molecular markers are effectively being used to detect the variant alleles present in diverse genotypes or cultivars which are phenotypically different from each other for the traits in question. Microsatellite markers or SSR markers are PCR-based markers that specifically bind to the complementary sequence of the plant genomic DNA during PCR reaction. Through allele mining novel genes could be identified from diverse sources and used in improvement of crops for specific traits. Allele mining for grain yield related traits in large number of genotypes/cultivars would help in identifying new alleles of the genes known for the target trait and their sources. It can also provide an insight into molecular basis of novel trait variations and identify the nucleotide sequence changes associated with superior alleles.
The quantitative trait locus (QTL) mapping has contributed to a better understanding of the genetic basis of many agronomically important traits such as grain yield. Grain yield is a quantitatively inherited trait which is the result collectively contributed by several individual grain traits. Of these, grain length, grain width and grain weight are some of the important yield contributing grain traits in rice. Today, a number of genes/QTLs associated with these traits have been identified, sequenced, mapped and many associated markers to these QTLs have been developed recently. In this study, a set of 41 diverse genotypes including two accessions viz. RWR- 19 and RWR-125 from wild species O. nivarawere used for allele mining using nine SSR markers tagged to reported genes/QTLs. Out of the nine primers viz. RDD 1-2, RM 234, RM 478, RM 23201, RM 502, RM 574-1, RM 574-2, RM 593-1 and RM 593-2 used in the present study, RM 23201 was found to be monomorphic and the rest eight primers were polymorphic.
Association between genotype and phenotype
Chi square test was conducted to study the association of the markers with grain traits. Each of the alleles observed in the top 15 and bottom 15 genotypes were tested for the consistency of their distribution with the whole population. Significant association (Χ2=3.94, p=0.05) was observed for allele A in the 15 genotypes having large grain length for primer RDD 1-2. Allele Bfor Rdd 1 gene too showed significant association (Χ2=4.51, p=0.03). Allele A was found to be associated with large grain length and allele B was found to be associated with small grain length. Significant association was observed for marker RM 478 with grain weight and for the distribution of A allele in the bottom 15 genotypes (lowest grain weight) with the distribution in the overall population (Χ2=5.08, p=0.02) and for primer RM 574-2, distribution of B allele (Χ2=4.3, p=0.08) in the bottom 15 genotypes (smallest grain width) was found to be significant. Rdd 1(Rice dof daily fluctuations 1) Dof gene in rice has been reported to be associated with plant growth, grain size and flowering time in rice. In a study conducted by Iwamoto et al. (2009), transgenic plants carrying full length Rdd 1 cDNA driven by constitutive promoter were produced in order to study the role of Rdd 1 and in vivo function of Dof gene in plants. The two types of transgenic plants were produced viz. RDD 1-S, transgenic containing sense strand and RDD 1-AS with anti-sense strand. Differences in the grain length, grain width and grain weight were observed between RDD 1-AS and RDD 1-S plants when subjected under varying photoperiod conditions [4]. When the grain sizes of the two transgenic types of plants were compared with the wild type plants, there was significant decrease in grain length, grain width and decrease in 1000 grain weight in transgenics carrying antisense gene (AS1=4.67 and AS2=4.54).
In our study, we used previously reverse RDD-1 primer and designed a new forward primer giving an amplicon size of 620bp. The three alleles obtained in the present study with sizes of 600, 620 and 600’490 base pairs were designated as A, B and B’C, respectively. Significant association (Χ2=3.94, p< 0.05) was observed between the grain length and the allele A (600bp) for large grain length and allele B (620bp) associated with small grain length (Χ2=4.51, p<0.05). The allele A for Rdd 1 can be used in breeding for large grain length. In our study, however, a set of 41 genotypes having large grain length ranging from 8.27-10.31 mm and small grain length of 5.48-7.52 mm were used. This phenotypic variation is distinct from the WT genotype Nipponbare (4.94mm) used by Iwamoto [4]. The allele A though similar in size could be different due to nucleotide variation. In order to rule out that possibility, sequencing of the allele from a panel of genotypes showing large grain length will have to be done. GS 5 (Grain size 5) QTL has been reported to be a positive regulator of grain width and flanked by markers RM 593 and RM 574. Over-expression of GS 5 promotes cell division resulting in an increase of grain width [6]. Other genes/QTLs in rice are Grain weight 7 (gw 7) reported to be associated with 1000 grain weight located at chromosome 7 in population derived from O. sativa and O. grandiglumis. Phenotypic variation of 13.3% was observed in populations having allele for gw 7 from Caipo [1]. Grain width 8 (gw 8) at chromosome 8 has been reported to have negative effect on grain width with decrease in grain width in populations resulting from separate crosses made between HJX 74 x Anmol 3 and HJX 74 x Basmati 370, with possible introgression of the recessive allele from Anmol 3 and Basmati 370 [7,8]. No significant association was observed for primers reported for Grain weight 8.1 (gw 8.1) [9] QTL in our panel of 41 genotypes.
Grain thickness 7 (gt 7), a QTL for grain thickness with linked marker RM 478 at chromosome 7¸ has been reported to be associated positively with grain thickness in HG 101 which is a near isogenic line obtained from O. sativa (Hwaseongbyeo) x O. grandiglumis [8]. The increased grain length in mm (5.53), grain width in mm (3.08) and 1000 grain weight in grams (26.3) was observed for the NILs as compared to the recipient parent, Hwaseongbyeo, with grain length (5.03mm), grain width (2.84mm) and 1000 grain weight (21.5g). In the present study, significant association of primer RM 478 was observed and the allele A (205bp) was significantly associated with low 1000 grain weight. The genotypes used in the present study have highest 1000 grain weight ranging from 20.16-36.04 grams and lowest ranging from 10.72-15.59 grams. Since grain yield is quantitative in nature, it is a complex trait and the markers tagged with the genes/QTLs may not show association in a diverse set of genotypes. Also the effect of individual genic components to grain yield may be very low as grain yield is a trait caused by cumulative action of many related components. Our study however, suggests that the allele A (600bp) from Rdd 1 can be used in breeding programmes to enhance grain length in rice. The genotypes viz. Hans raj, Pant dhan-18, Pant dhan- 10, Narendra, Pant dhan-12, Pant dhan-4, ARR-09, Shahsarang, IVTASG- 2712, IVT-ASG-2701, AVT-IASG-2602, AVT-IASG-2609 and IVTASG- 2705 have been identified to be carrying this allele. These can be used as a source of large grain length in future. The primers RM 478 for grain thickness and RM 574-2 for grain width can be used in marker assisted breeding programmes. Allele A (205bp) of RM 478 and allele B (240bp) of RM 574-2 associated with low grain weight and grain width, respectively can be used for rejection at seedling stage in marker assisted breeding programme. This study also identified novel alleles (alleles distinct from those reported in biparental programme); which could be significant in a larger panel of rice genotypes (Tables 7-9).
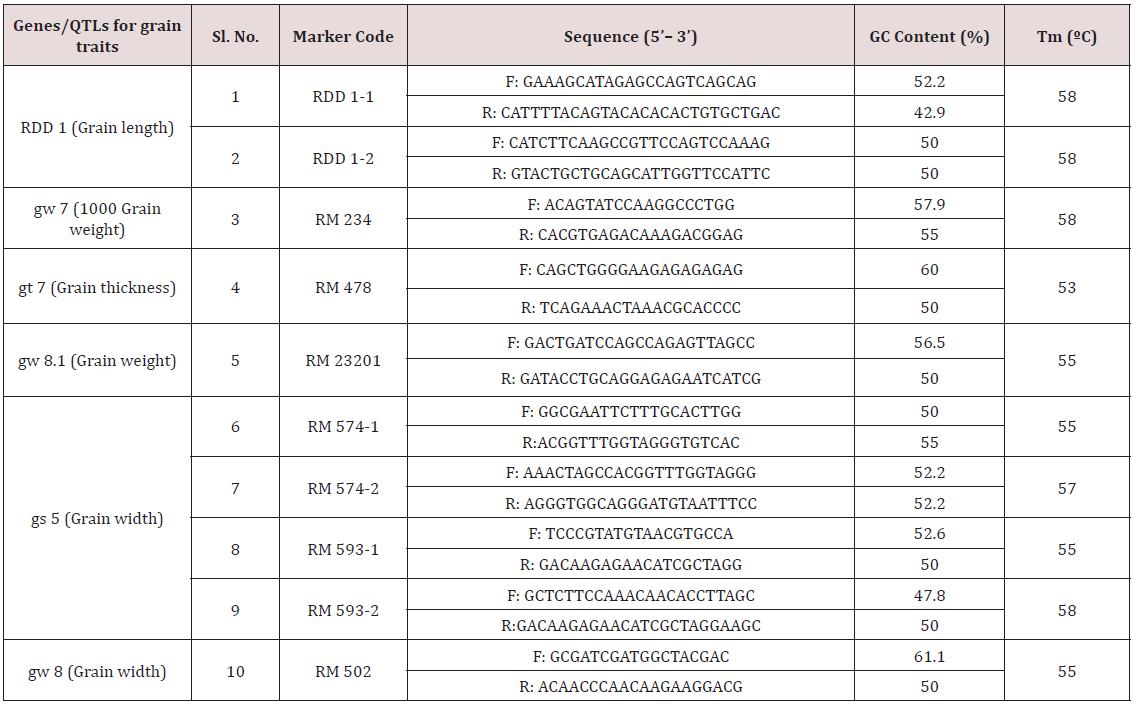
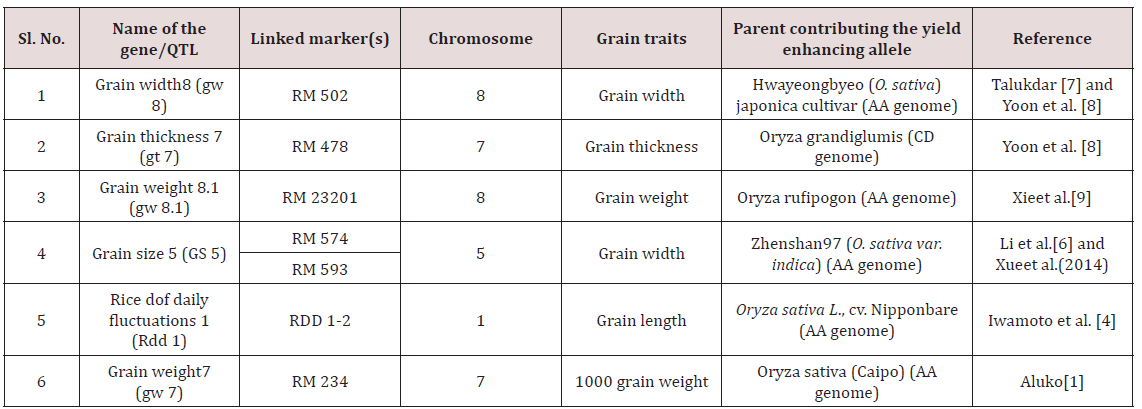
Table 7: List of reported candidate genes/QTL, their associated markers, location, and associated grain traits used for allele mining study.
Conclusion
From this present experiment, hybrid plants from inter-specific crosses can be used for backcrossing in future plant breeding programmes. Crosses can also be attempted for genotypes that failed during this experiment. Primer RDD 1-2 can be used for identifying alleles enhancing grain length from diverse genomic sources for crop improvement. The allele A (600 bp) from Rdd 1 can be used in breeding programmes to enhance grain length in rice. The genotypes viz. Hans raj, Pant dhan-18, Pant dhan-10, Narendra, Pant dhan-12, Pant dhan-4, ARR-09, Shahsarang, IVTASG- 2712, IVT-ASG-2701, AVT-IASG-2602, AVT-IASG-2609 and IVT-ASG-2705 identified carrying this allele can be used as a source for large grain length in future. The primers RM 478 for grain thickness and RM 574-2 for grain width can be used in marker assisted breeding programmes. Allele A (205bp) of RM 478 and allele B (240bp) of RM 574-2 associated with low grain weight and grain width respectively, can be used for rejection at seedling stage in marker assisted breeding programme. This study also identified novel alleles (alleles distinct from those reported in biparental programme); which could be significant in a larger panel of rice genotypes.
References
- Aluko GK (2003) Genetic mapping of agronomic traits from the interspecific cross Oryza sativa (L.) and Oryza glaberrima (Steud.). PhD. Dissertation, submitted to the Graduate Faculty of the Louisiana State University and Agriculture and Medical College, USA.
- Amante Bordeous AD, Sitch LA, Nelson RJ, Dalmaclo RD, Oliva NP, et al. (1992) Transfer of bacterial blight and blast resistance from the tetraploid wild rice Oryza minutato cultivated rice, Oryza sativa. Theor Appl Genet 84(3-4): 345-354.
- Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissues. Focus 12(1): 13-15.
- Iwamoto M, Higo K, Takano M (2009) Circadian clock and phytochrome regulated Dof-like gene, Rdd 1, is associated with grain size in rice. Plant, Cell and Environment 32(5): 592-603.
- Li J, Xiao J, Grandillo S, Jiang L, Wan Y, et al. (2004) QTL detection for rice grain quality traits using an inter-specific backcross population derived from cultivated Asian (O. sativa L.) and African (O. glaberrima S.) rice. Genome 47(4): 697-704.
- Li Y, Fan C, Xing Y, Jiang Y, Luo L, et al. (2011) Natural variation in GS 5 plays an important role in regulating grain size and yield in rice. Nat Genet 43(12): 1266-1269.
- Talukdar A, Zhang GQ (2008) Inheritance of grain shape and molecular tagging of the QTL for reduced grain width (gw) in rice (Oryza sativa L.). Indian J Genet 68(2): 103-107.
- Yoon DB, Kang KH, Kim HJ, Ju HG, Kwon SJ, et al. (2006) Mapping quantitative trait loci for yield components and morphological traits in an advanced backcross population between Oryza grandiglumis and the Oryza sativa japonica cultivar Hwaseongbyeo. Theor Appl Genet 112(6): 1052-1062.
- Xie X, Song MH, Jin F, Ahn SN, Suh JP, et al. (2006) Fine mapping of a grain weight quantitative trait locus on rice chromosome 8 using nearisogenic lines derived from a cross between Oryza sativa and Oryza rufipogon. Theor Appl Genet 113(5): 885-894.

Top Editors
-

Mark E Smith
Bio chemistry
University of Texas Medical Branch, USA -

Lawrence A Presley
Department of Criminal Justice
Liberty University, USA -

Thomas W Miller
Department of Psychiatry
University of Kentucky, USA -

Gjumrakch Aliev
Department of Medicine
Gally International Biomedical Research & Consulting LLC, USA -

Christopher Bryant
Department of Urbanisation and Agricultural
Montreal university, USA -

Robert William Frare
Oral & Maxillofacial Pathology
New York University, USA -

Rudolph Modesto Navari
Gastroenterology and Hepatology
University of Alabama, UK -

Andrew Hague
Department of Medicine
Universities of Bradford, UK -

George Gregory Buttigieg
Maltese College of Obstetrics and Gynaecology, Europe -

Chen-Hsiung Yeh
Oncology
Circulogene Theranostics, England -
.png)
Emilio Bucio-Carrillo
Radiation Chemistry
National University of Mexico, USA -
.jpg)
Casey J Grenier
Analytical Chemistry
Wentworth Institute of Technology, USA -
Hany Atalah
Minimally Invasive Surgery
Mercer University school of Medicine, USA -

Abu-Hussein Muhamad
Pediatric Dentistry
University of Athens , Greece

The annual scholar awards from Lupine Publishers honor a selected number Read More...