
Lupine Publishers Group
Lupine Publishers
Menu
ISSN: 2644-1209
Research ArticleOpen Access
Extracellular Proteases from Keratitis Causing Fusarium, Aspergillus and Dematiaceous Species Volume 2 - Issue 2
Swati N Madhu1, Anuradha K Pal2 and Devarshi U Gajjar1*
- 1 Department of Microbiology and Biotechnology Centre, Gujarat, India
- 2 Iladevi Cataract and IOL Research Centre, Karnataka, India
Received:January 30, 2019; Published: February 08, 2019
Corresponding author: Devarshi Gajjar, Department of Microbiology and Biotechnology Centre, Gujarat, India
DOI: 10.32474/TOOAJ.2019.02.000132
Abstract
Purpose
Various types of hydrolytic enzymes, like proteases, lipases and phospholipases are important virulence factors produced by pathogenic fungi. In the present study quantification and characterization of proteases produced by different pathogenic Aspergillus, Fusarium and Dematiaceous Species isolated from corneal ulcers was done.
Method
Seven Aspergillus species, twelve Fusarium species and five Dematiaceous Species previously isolated from corneal ulcers were used in the present study. All isolates were grown in Sabarouds Dextrose Broth (SDB) and culture filtrate was obtained after 10 days of growth. Acetone precipitated culture filtrate was used as the source of protease. Characterization of proteases was done by assaying the activity at different pH (6-12), with inhibitors (PMSF, Pepstatin and EDTA) and gelatin zymography.
Result
In Aspergillus spp, maximum specific activity of protease was found in Aspergillus flavus and minimum specific activity was found in Aspergillus niger and Aspergillus tubingensis. In Fusarium spp., proteases from Fusarium solani Species complex (FSSC) showed more activity compared to F. delphinoides. Proteases from Aspergillus spp. and FSSC showed inhibition in presence of PMSF, while EDTA inhibited the proteases from F. delphinoides. In Dematiaceous group, Curvularia spp showed highest activity and were inhibited by PMSF.
Conclusion
It was concluded that fungi causing keratitis produce varying amounts and types of proteases. The difference in the specific activity of extracellular proteases among the isolates could indicate the differences in their pathogenic potential.
Keywords:Mycotic keratitis; Aspergillus spp; Fusarium spp; Proteases; Virulence factors
Introduction
Corneal blindness, in developing countries is predominantly associated with infections. In India, nearly 35-50% of all infectious keratitis are caused by fungi [1]. The term mycotic keratitis refers to a corneal infection caused by fungi. More than 105 species of fungi belonging to 56 genera have been reported to cause fungal keratitis. The most common isolates which are isolated from the eyes of the patients are species of Fusarium, Aspergillus, Candida and other hyaline and Dematiaceous hyphomycetes. The most common risk factor for keratitis is ocular trauma through vegetative material while, other predisposing factors can be prolonged use of topical corticosteroids or antimicrobial agents, systemic diseases such as diabetes mellitus, preexisting ocular diseases and use of contact lenses.
Pathogenic fungi retain several factors which allow their growth in adverse conditions provided by the host and contribute to disease development. These factors are known as virulence factors which favor the infection process and are important determinants of the pathogenic potential of any organism [2]. The pathogenecity of the fungus may be due to fungal cell wall components, toxins, enzymes or pigments produced by fungus. The microbial attributes that confer the potential for virulence fall primarily within several categories, including the ability to enter a host; the ability to evade host defenses; to grow in a host environment; to counteract host immune responses; to acquire iron and nutrients from the environment and to sense environmental change. Production and secretion of hydrolytic enzymes, such as proteases, lipases and phospholipases are among the important virulence traits [3].
Proteases, also termed as proteinases or peptidases, are proteolytic enzymes which function as molecular knives which cut long amino acid sequences into fragments, a process that is essential for the synthesis of all proteins, controlling their size,composition, shape, turnover, and ultimate destruction[4]. These enzymes play a role in nutrition, tissue damage, dissemination within the human organism, iron acquisition and overcoming the host immune system and hence strongly contribute to fungal pathogenicity [3]. Extracellular proteases from Fusarium spp. and Aspergillus spp. cause corneal destruction in experimental rabbit models [2,5]. Proteolytic activities of A. flavus and F. solani in rabbit corneas during active fungal keratitis were investigated in vivo and it was found that the fungal cultures predominantly produced serine proteases in vitro; while, fungal infected corneal homogenates showed the presence of metalloproteases alone [5]. Production of proteases in Aspergillus and Fusarium causing keratitis has also been shown to be correlating significantly with amphotericin B resistance [6]. Further, alkaline protease (Alp1) has also been shown to be one of the abundant proteins in A. flavus exoproteome [7]. Taken together, these studies indicate that proteases can mediate corneal invasion and could also cause corneal melting. However, there is paucity of information regarding their detailed characterization from keratitis causing isolates. Earlier we reported a prospective study to compare different aspects of fungal keratitis such as its clinical features, microbial evaluation, molecular identification, antifungal susceptibility, and clinical outcomes between Fusarium, Aspergillus and Dematiaceous fungi [8]. In the present study detailed analysis of proteases produced by these three groups of fungi is attempted.
Material and Methods
Fungal Isolates
Clinical isolates of fungi (n=24); belonging to Fusarium spp. (n=12); Aspergillus spp. (n=7) and Dematiaceous spp. (n=5) isolated in the previous study and were available in the lab [8,9]. All isolates were identified using the ITS region sequencing and sequences were deposited in NCBI. Name and GenBank accession no of fungi (n=25) used in the present study are: Fusarium delphinoides (n=4) [Cc26- KM017139, Cc52- JQ910153, Cc 119- KM017141, Cc132- KM017140]; Fusarium solani Species complex (FSSC) (n=8) [Cc50-KM017134, Cc149- JQ910159, Cc163- KM017142, Cc167-KM017143, Cc172-KM017144, Cc204-KM017135, Cc215- KM017137, Cc256-KM017138]; Aspergillus flavus (n=1) [Cc59- JQ910160]; Aspergillus sydowii (n=1) [Cc73-JQ910152]; Aspergillus niger (n=1) [Cc101- JQ910154]; Aspergillus terreus (n=1) [Cc112- KM017145]; Aspergillus tubingensis (n=1) [Cc117-JQ910156]; Aspergillus tamarii (n=1) [Cc129- JQ910158]; Aspergillus fumigatus (n=1) [Cc249- KM017131]; Curvularia lunata (n=3) [Cc70- KM017133, Cc90-JQ910155, Cc157-KM017132]; Phomopsis phoenicicola (n=1) [Cc58- HQ650813]; Phaeoacremonium rubrigenum (n=1) [Cc79-KM017136].
Growth of Fungi
The isolates were first grown on Sabouraud’s Dextrose Agar (SDA) (Himedia Laboratories, India) and after 7 days of growth, approximately 10 mm disc was cut from the SDA plate and inoculated in 100 ml of Sabouraud’s Dextrose Broth (SDB) in 500 ml conical flask. The fungus was grown on SDB for 10 days at 30±2 ºC in static condition.
Extraction and Precipitation of Proteases
After 10 days of fungal growth, the culture filtrate was collected using sterile Whatman filter paper in a sterile bottle. The obtained culture filtrate was then subjected to precipitation using chilled acetone. To 1 ml of fungal culture filtrate 4 ml of chilled acetone was added drop wise. Culture filtrate – acetone mixture was then kept at -26 ºC over night for precipitation. Next day, the precipitate was collected through centrifugation at 5000 RPM for 15 minutes at 4 ºC. Resultant precipitate was dissolved in Phosphate buffered Saline (PBS) (50 μl PBS/ml of initial culture filtrate used for precipitation) and used for azocaesin assay and gelatin zymography. Protein estimation of concentrated extract was done by the method of Lowry et al. [10].
Specific Activity Measurement by Azocasein Assay
Azocasein (Sigma Aldrich, USA) was dissolved at a concentration of 5 mg/ml in assay buffer containing 50 mM Tris (Sigma Aldrich, USA) (pH 7.4), 0.2M NaCl (Himedia Laboratories, India), 5mM CaCl2 (Himedia Laboratories, India), 0.05% Brij 35 (Himedia Laboratories, India), and 0.01% sodium azide (Himedia Laboratories, India). The azocasein solution (400μl) was mixed with 100μl of precipitated culture supernatants and incubated in a 37°C water bath for 90 min. The reactions were stopped by adding 150μl of 20% trichloroacetic acid (Sigma Aldrich, USA), and the reaction mixtures were allowed to stand at the ambient temperature for 30 min. Tubes were then centrifuged for 3 min at 8,000g, and 50μl of each supernatant was added to 500μl of 1 M NaOH (Himedia Laboratories, India). The absorbance at 436nm of released azo dye was determined. One unit of enzyme activity was defined as an increase of 0.1 absorption unit after incubation for 1 hour. Specific activity was calculated as Units/ mg.
Characterization of Proteases
For characterization of proteases; the effect of pH and inhibitors was studied using the precipitated enzyme extracted from all fungi. Effect of pH: The effect of pH on enzyme activity was studied by incubating the enzyme in azocasein solution of respective pH. Azocasein activity was studied at 6 different pH ranging from acidic to basic i.e. pH 3, pH 5, pH 7.4, pH 8, pH 10 and pH 12. Effect of enzyme inhibitors: To understand the type of enzyme produced, specific enzyme inhibitors were used and azocasein assay was carried out. PMSF (Sigma Aldrich, USA) (inhibitor of serine protease at final concentration of 0.5mM), EDTA (Himedia Laboratories, India) (inhibitor of metallo protease at final concentration of 1mM) and Pepstatin (Sigma Aldrich, USA) (inhibitor of aspartyl proteases at final concentration 1μM) was used to check the type of proteases present in fungal isolates. Proteases samples were incubated with inhibitor for 1 hour and same concentration of inhibitor was also added in azocasein solution in which the azocasein assay was carried out. Azocasein assay was done by the method described earlier.
Gelatin Zymography
Zymography was performed using gelatin (Sigma Aldrich, USA) as a substrate (0.1%) and 12% of polyacrylamide (Sigma Aldrich, USA). Twelve percent SDS- polyacrylamide gel was prepared with 0.1% gelatin. Enzyme units (0.2) were taken, mixed with 6X loading dye and electrophoresed. After electrophoresis, the gel was incubated in incubation buffer (50mM Tris, 5 mM CaCl2 and 1 μM ZnCl2) containing 2.5 % Triton –X 100 (Himedia Laboratories, India) for 1 hour. The gel was washed twice with deionized water and again kept in same buffer at 37°C for 2 hours. The gel was again washed with deionized water and kept overnight for staining with (0.5%) Commassie Brilliant blue R (Himedia Laboratories, India). The gel was de-stained with destaining solution (methanol: water: acetic acid, 4:5:1v/v) to observe the bands of enzyme activity. The bands of substrate degradation were observed as colorless bands against blue background. Zymography in presence of inhibitors was also done. The proteases samples were pre-incubated with desired concentration of three inhibitors i.e. PMSF, pepstatin and EDTA for 1 hour and then loaded into gel for zymography.
Results
Table 1: Comparative chart showing the activity of protease enzyme extracted from different fungus at different pH.
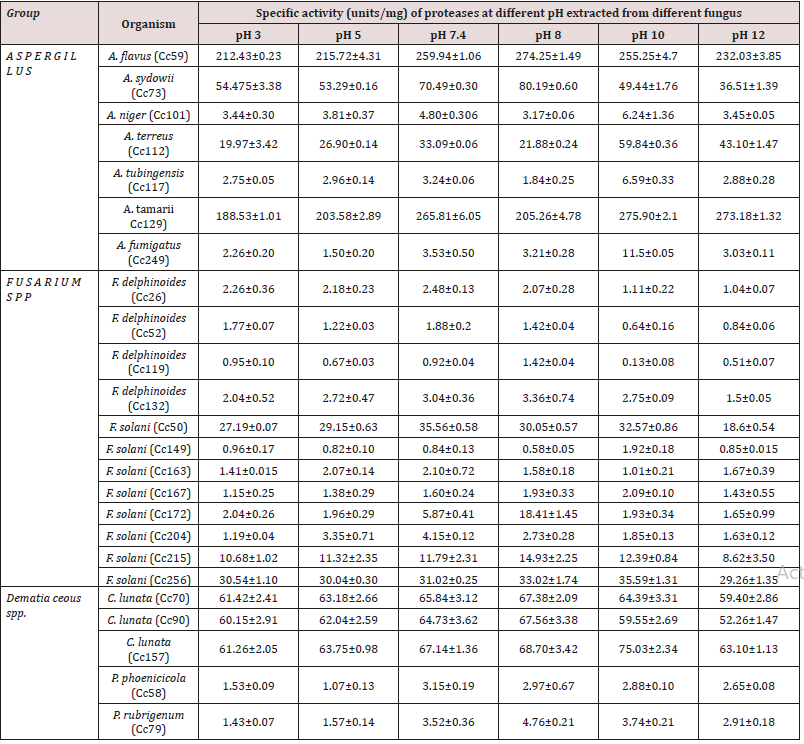
Table 1 shows the specific activity of extracellular proteases from all 24 fungi measured at 6 different pH values (pH 3, pH 5, pH 7.4, pH 8, pH 10 and pH 12).
Aspergillus Proteases
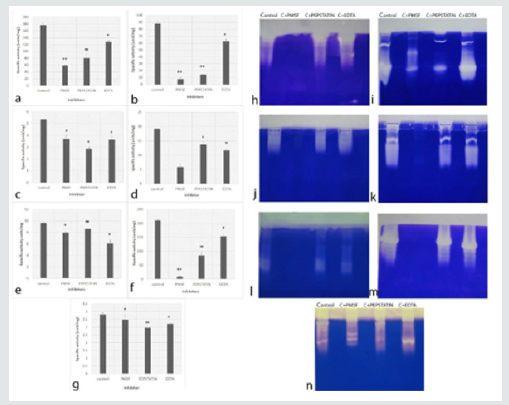
Proteases produced by Aspergillus species showed activity at all pH tested (Table1). The optimum pH of proteases from A. niger, A. terreus, A. tubingensis and A. fumigatus was pH 10. Proteases from A. flavus and A. sydowii showed high activity at pH 8; whereas, A. tamarii presented the activity of proteases at broad pH range. Out of 7 Aspergillus species used in the present study, A. flavus and A. tamarii illustrated more activity i.e. 274.25±1.49 units/mg and 275.9±2.1 units/mg, respectively as compared to rest of the species. A. niger and A. tubingensis showed relatively less specific activity which was 6.24±1.36 units/mg and 6.59±0.33 units/mg respectively at pH 10. The specific activity of A. sydowii was 80.19±0.60 units/mg at pH 8. In A. fumigatus, and A. terreus the specific activity was 11.5±0.05 units/mg and 59.84±0.36 units/mg respectively at pH 10. The results showed that the proteases produced by all Aspergillus species were mainly alkaline proteases. The in vitro inhibition assays showed that proteases from all 7 species were inhibited in presence of all 3 inhibitors with a significant difference with p value at least < 0.05. The proteases inhibition pattern in tube assay and zymography gels for A. flavus and A. tamarii was similar, where the maximum inhibition was observed in presence of PMSF followed by pepstatin and EDTA (Figures 1a & f & h & m). In A. terreus, the inhibition was highest in PMSF followed by EDTA and Pepstatin in tube assay (Figure 1d). However, in zymography, inhibition was only in presence of PMSF (Figure 1k). The proteases from A. fumigatus showed maximum inhibition by pepstatin in both in vitro assay and zymography gel (Figures 1g & 1n). However, in A.niger pepstatin showed highest inhibition in tube assay while, PMSF showed maximum inhibition in zymography gel (Figures 1c & 1j). Similar discrepancy was seen in A. tubingensis, the proteases were inhibited maximum in presence of EDTA in tube assay but were inhibited by PMSF in zymography gel (Figures 1e & 1l). These results suggest that the type of proteases produced by Aspergillus is mainly serine proteases, followed by aspartyl and metallo proteases.
Figure 1: Effect of inhibitors (PMSF, Pepstatin and EDTA) on Aspergillus proteases a: A. flavus (Cc59), b: A. sydowii (Cc73), c: A. niger (Cc101), d: A. terreus (Cc112), e: A. tubingensis (Cc117), f: A. tamarii (Cc129), g: A. fumigatus (Cc249) (ns - Non significant, # - p value <0.05, ## - p value <0.02, * - p value <0.01 and * * - p value < 0.001). h-n: 12% Zymography gel using gelatin (0.1%) as a substrate of A. flavus (Cc59), A. sydowii (Cc73), A. niger (Cc101), A. terreus (Cc112), A. tubingensis (Cc117), A. tamarii (Cc129), A. fumigatus (Cc249), respectively.
Fusarium Proteases
In Fusarium, the proteases from F. delphinoides showed activity at broad pH range between pH 3 to pH 8 (Table 1). The specific activity of proteases for different F. delphinoides isolates was ranging between 1.42±0.04 units/mg in Cc 119 to 3.36±0.74 units/mg in Cc132. In case of FSSC the value of specific activity was highly variable among species and the proteases were active at broad pH range. Isolate no Cc50 and Cc256 of FSSC showed highest specific activity of 35.56±0.58 units/mg and 35.59 ± 1.31 units/mg respectively at alkaline pH range (pH 7.4 to pH 10) and the lowest specific activity was observed in Cc149 (1.92 ± 0.18 units/mg) at pH 10. The results showed that the proteases in F. delphinoides were acidic to neutral, and the proteases from FSSC were variable in alkaline range.
Figure 2: Effect of inhibitors (PMSF, Pepstatin and EDTA) on F. delphinoides proteases a: (Cc26), b: (cc52), c: (Cc119), d: (Cc132), (ns- non significant, * - p value <0.01 and ** - p value < 0.001), e: Representative 12% Zymography gel using gelatin (0.1%) as a substrate. No inhibition was observed in presence of any of the inhibitor.
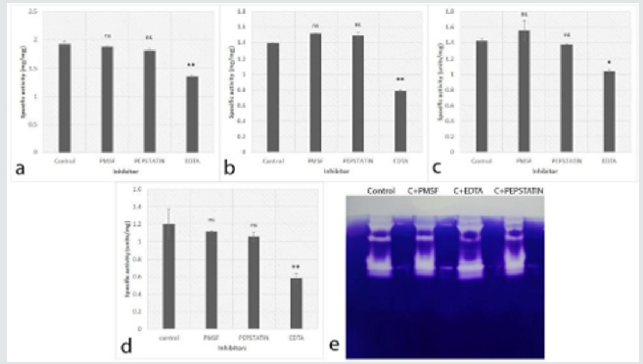
Figure 3: Effect of inhibitors (PMSF, Pepstatin and EDTA) on F. solani proteases. a: (Cc50), b: (Cc149), c: (Cc163), d: (Cc167), e: (Cc172), f: (Cc204), g: (Cc215), h: (Cc256) (ns- non significant, # - p value <0.05, ## - p value <0.02* - p value <0.01 and ** - p value < 0.001) i: Representative 12% Zymography gel using gelatin (0.1%) as a substrate. The inhibition of proteases was observed in the presence of PMSF.
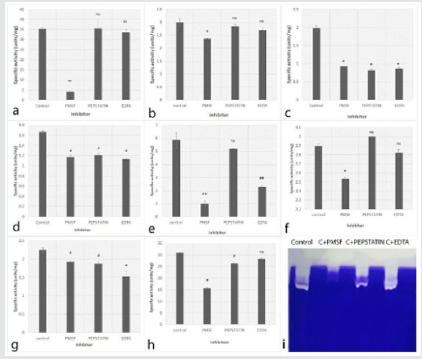
The proteases extracted from all four F. delphinoides showed inhibition in presence of EDTA in both tube assay and zymography (Figure 2). The results indicate that the proteases present in F. delphinoides are mainly metallo proteases. All eight FSSC proteases were inhibited in presence of PMSF (Figure 3). In case of Cc163, Cc167, Cc 215 and Cc 256 the inhibition was noticed in presence of all the three inhibitors i.e. PMSF, Pepstatin and EDTA. However, in rest of the four species, the inhibition was found only in presence of PMSF, which showed that the majority of proteases in Cc50, Cc149, Cc 172 and Cc204 are serine proteases. Zymography indicated that, the proteases from FSSC showed the inhibition in presence of PMSF only, and no difference was found with other inhibitors (Figure 3i).
Dematiaceous Proteases
In Dematiaceous group of fungi, the proteases were studied in three Curvularia lunata, one Phomopsis phoenicicola and one Phaeoacremonium rubrigenum. Proteases from all three C.lunata isolates had activity at all pH, with slight more activity at pH 8 and pH 10 (Table 1). The maximum activity of proteases was illustrated by Curvularia isolates Cc 70, Cc 90 and Cc 157 was 67.38±2.09 units/ mg, 67.56±3.38 units/mg and 75.03±2.34 units/mg respectively. Proteases obtained from P.phoenicicola showed high activity at pH 7.4 with the specific activity of 3.15±0.19 units/mg (however, it had activity at broad pH range) and P. rubrigenum proteases presented maximum activity (4.76±0.21 units/mg) at pH 8. The tube inhibition assays from C. lunata isolates (Cc90 and Cc157) showed inhibition in presence of PMSF, followed by pepstatin and no inhibition in presence of EDTA (Figure 4). However, zymography gels showed inhibition only by PMSF (Figure 4d). In tube inhibition assays for proteases from P. phoenicicola and P.rubrigenum, inhibition was noted by pepstatin and EDTA respectively. However, slight inhibition was observed in zymography gels (Figures 4g & h). The results of the specific activity of proteases produced by 24 different isolates suggests that; the number of extracellular proteases was more in Aspergillus and Curvularia species compared to Fusarium species. The overall summary of proteases from different isolates is given in Table 2.
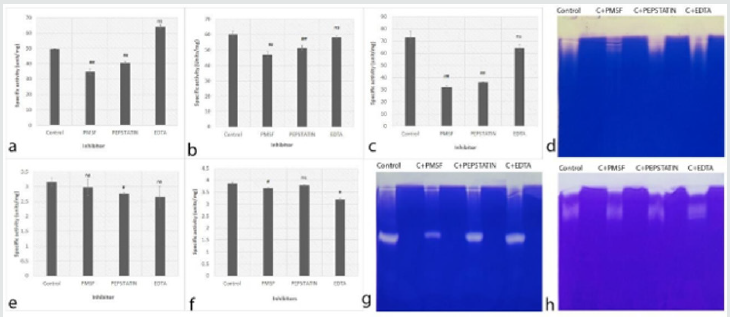
Table 2: Summary of proteases for optimum pH, effective inhibitors and type of proteases present in Aspergillus species, Fusarium species and dematiaceous fungi.
Figure 4: Effect of inhibitors (PMSF, Pepstatin and EDTA) on Dematiaceous fungi proteases. a: C. lunata (Cc90), b: Cladosporium spp (Cc127), c: C. lunata (Cc157) e: Phomopsis phoenicicola (Cc58), f: Phaeacremonium (cc79) (ns- non significant, # - p value <0.05, # # - p value <0.02* - p value <0.01 and ** - p value <0.001), d,g,h: Representative 12% Zymography gel using gelatin (0.1%) as a substrate of culvularia spp., P. phoenicicola and P. rubrigenum, respectively.
Discussion
In the present study, extracellular protease activity was quantified from three most common keratitis causing fungal pathogens; Aspergillus spp., Fusarium spp., and Dematiaceous fungi. Among the three, Aspergillus isolates produced the highest amount proteases while, Fusarium and Dematiatous produced moderate amounts. Proteases from Aspergillus spp. causing keratitis to have been reported only from Aspergillus flavus [5,11,12]. The exoproteome analysis of a keratitis causing A. flavus showed that nearly 50% of the exoproteins possess catalytic activity and one of these, an alkaline serine protease (Alp1) is abundant and present in multiple proteoforms [7]. Proteases from none of the other species has been studied from keratitis causing isolates. We have quantified proteases from A. sydowii, A. niger, A. terreus, A. tubingensis, A. tamarii and A. fumigatus. Proteases from soil isolates of Aspergillus are extensively studied and described from A. niger [13], A. tamarii [14], A. fumigatus [15], A. flavus, A. terreus [16] and A. tubingensis [17]. Total extracellular protease activity for some isolates (A. sydowii, A. tubingensis, A.terreus) was higher than the reported isolates; while, it was less for certain isolates (A. flavus, A niger, A. tamarii). However, these differences could be because of the differences in the isolates used and conditions for their growth. Large variations in occurrence and abundance of proteases in seven Aspergillus spp. was shown using genome mining and comparative proteomics [18]. Serine proteases were the largest group in the protease spectrum across Aspergillus spp. The alkaline nature of extracellular proteases from all Aspergillus spp. in the present work corroborates with most of the published reports.
Extracellular enzymes such as lipase, deoxyribonuclease (DNase), α-amylase, protease, cellulase and pectinase produced by the Fusarium isolates from keratitis were screened on solid media supplemented with the corresponding substrates [19]. The authors showed significant differences in protease activity between clinical and soil isolates. Fusarium proteases have been documented in rabbit model for keratitis [5]. However, detailed characterization regarding the types, number and amount of proteases are lacking from keratitis causing Fusarium isolates. In the present work, preliminary characterization of proteases from four F. delphinoides and eight F. solani isolates was done. Our results showed the presence of extracellular metalloprotease in F. delphinoides. In two F. solani isolates (CC50 and CC149), serine proteases were found while, the rest showed the presence of serine, aspartyl and metallo protease. To the best of literature search, there are no reports of proteases in F. delphinoides. Proteases have been reported earlier from soil and plant pathogenic isolates; F. solani [20], F. oxysporum [21] and F. venenatum [22]. Hence, species of Fusarium are known to produce different types and amounts of extracellular proteases. Most of these studies report the presence of one major serine protease in Fusarium spp. Zymography results in the present study indicate the presence of more than one type of high molecular weight protease in Fusarium. These results warrant further detailed studies on purification and identification of these proteases.
In Dematiaceous group, our results indicate the presence of serine and aspartyl proteases in Curvularia spp, aspartyl proteases in P. phoenicicola and serine and metallo protease in P. rubrigenum. Serine protease from C.lunata has been reported [23], but presence of aspartyl proteases was not found. Molecular and immunological characterization of subtilisin like serine proteases was carried out in C. lunata, and the protease was proved as a major allergen responsible for inflammation and asthma [24]. To the best of our knowledge there are no studies on extracellular proteases in P. phoenicicola species, but P. azadirachtae has been explored for the production of extracellular enzymes like polygalacturonase, laccase, protease and xylanase [25].
Our study showed that serine proteases were the major proteases found in isolates of all the groups (Aspergillus, Fusarium and Dematiaceous) of fungi. Among the three groups; Aspergillus spp. had the highest level of proteases, followed by Fusarium and Dematiaceous group. These results corroborate with our clinical findings of patients from which these fungi were isolated [9]. Aspergillus had the maximum number of worsened cases; followed by Fusarium and then Dematiaceous group; suggesting that Aspergillus spp. are more virulent compared to the other two [9]. The serine proteinases and can degrade elastin, collagen, fibrin and fibrinogen. Many studies have correlated the extracellular proteases production with virulence of fungi [26-28]. Hence, collagenase activity may be a mediator of the severe corneal destruction. Recently, we showed the presence of extracellular serine proteases using the ex vivo goat cornea model using Fusarium as the pathogen [29-31]. Further studies to identify the proteases in ongoing.
Conclusion
To conclude, fungal isolates showed the presence of various types of proteases in varying amounts. Proteolytic activity leads to severe corneal destruction and hence, there is a need to identify these proteases. Identification of such proteases could help in identifying molecules which inhibit them. Future strategies should encourage the development of compounds to combat virulence mechanisms as potential targets against drug resistant organisms.
References
- Chidambaram JD, Venkatesh Prajna N, Srikanthi P, Lanjewar S, Shah M, et al. (2018) Epidemiology, risk factors, and clinical outcomes in severe microbial keratitis in South India. Ophthalmic Epidemiol 25(4): 297- 305.
- Brunke S, Mogavero S, Kasper L, Hube B (2016) Virulence factors in fungal pathogens of man. Curr Opin Microbiol 32: 89-95.
- Karkowska-Kuleta J, Rapala-Kozik M, Kozik A (2009) Fungi pathogenic to humans: Molecular bases of virulence of Candida albicans, Cryptococcus neoformans and Aspergillus fumigatus. Acta Biochim Pol 56(2): 211- 224.
- Shen HB, Chou KC (2009) Identification of proteases and their types. Anal Biochem 385(1): 153-160.
- Gopinathan U, Ramakrishna T, Willcox M, Rao CM, Balasubramanian D, Kulkarni A, et al. (2007) Enzymatic, clinical and histologic evaluation of corneal tissues in experimental fungal keratitis in rabbits. Exp Eye Res 72(4): 433-442.
- Nayak N, Satpathy G, Prasad S, Vajpayee RB, Pandey RM (2010) Correlation of proteinase production with amphotericin B resistance in fungi from mycotic keratitis. Ophthalmic Res 44(2): 113-118.
- Selvam RM, Nithya R, Devi PN, Shree RS, Nila MV, et al. (2015) Exoproteome of Aspergillus flavus corneal isolates and saprophytes: identification of proteoforms of an oversecreted alkaline protease. J Proteomics 115: 23-35.
- Gajjar DU, Pal AK, Ghodadra BK, Vasavada AR (2013) Microscopic evaluation, molecular identification, antifungal susceptibility, and clinical outcomes in Fusarium, Aspergillus and, Dematiaceous keratitis. Biomed Res Int 2013: 605308.
- Gajjar DU, Pal AK, Parmar TJ, Arora AI, Ganatra DA, Kayastha FB, et al. (2011) Fungal scleral keratitis caused by Phomopsis phoenicicola. J Clin Microbiol 49(6): 2365-2368.
- Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193(1): 265- 275.
- Leema G, Kaliamurthy J, Geraldine P, Thomas PA (2010) Keratitis due to Aspergillus flavus: clinical profile, molecular identification of fungal strains and detection of aflatoxin production. Mol Vis 16: 843-854.
- Zhu WS, Wojdyla K, Donlon K, Thomas PA, Eberle HI (1990) Extracellular proteases of Aspergillus flavus. Fungal keratitis, proteases, and pathogenesis. Diagn Microbiol Infect Dis 13(6): 491-497.
- Abdel Wahab WA, Ahmed SA (2018) Response surface methodology for production, characterization and application of solvent, salt and alkalitolerant alkaline protease from isolated fungal strain Aspergillus niger WA 2017. Int J Biol Macromol 115: 447-458.
- Anandan D, Marmer WN, Dudley RL (2007) Isolation, characterization and optimization of culture parameters for production of an alkaline protease isolated from Aspergillus tamarii. J Ind Microbiol Biotechnol 34(5): 339-347.
- Monod M, Togni G, Rahalison L, Frenk E (1991) Isolation and characterisation of an extracellular alkaline protease of Aspergillus fumigatus. J Med Microbiol 35(1): 23-28.
- Chellapandi P (2010) Production and Preliminary Characterization of Alkaline Protease from Aspergillus flavus and Aspergillus terreus. E-Journal of Chemistry 7: 479-482.
- Morya Y, Yadav D (2009) Production and partial characterization of neutral protease by an indigenously isolated strain of Aspergillus tubingensis NIICC-08155. Internet Journal of Microbiology 8(1): 53-55.
- Budak SO, Zhou M, Brouwer C, Ad Wiebenga, Isabelle Benoit, et al. (2014) A genomic survey of proteases in Aspergilli. BMC Genomics 15(1): 523.
- Mythili A, Babu Singh YR, Priya R, Shafeeq Hassan A, Manikandan P, et al. (2014) In vitro and comparative study on the extracellular enzyme activity of molds isolated from keratomycosis and soil. Int J Ophthalmol 7(5): 778-784.
- Bhuvaneswari P, Balasubramanian R (1999) Partial purification and properties of protease from Fusarium solani (Mart.) Sacc., a fungal associate of Puccinia archidis Speg. Indian J Exp Biol 37: 1113-1116.
- Barata RA, Andrade MH, Rodrigues RD, Castro IM (2002) Purification and characterization of an extracellular trypsin-like protease of Fusarium oxysporum var. lini. J Biosci Bioeng 94(4): 304-308.
- Farnworth NE, Robson GD, Trinci AP, Wiebe MG (2003) Trypsin-like protease (TLP) production in Fusarium oxysporum and Fusarium venenatum and use of the TLP promoter for recombinant protein (glucomylase) production. Enzym Microb Technol 33(1): 85-91.
- Gupta R, Sharma V, Sridhara S, Singh BP, Arora N (2004) Identification of serine protease as a major allergen of Curvularia lunata. Allergy 59(4): 421-427.
- Tripathi P, Nair S, Singh BP, Arora N (2011) Molecular and immunological characterization of subtilisin like serine protease, a major allergen of Curvularia lunata. Immunobiology 216(3): 402-408.
- Vedashree S, Sateesh MK, Lakshmeesha TR, Shahi Sofi M, Vedamurthy AK (2013) Screening and assay of extracellular enzymes in Phomopsis azadirachtae causing die-back disease of neem. Journal Agric Technol 9(4): 915-927.
- Burda CD, Fisher E (1960) Corneal destruction by extracts of Cephalosporium mycelium. Am J Ophthalmol 50: 926-937.
- Dudley MA, Chick EW (1964) Corneal Lesions Produced in Rabbits by an Extract of Fusarium Moniliforme. Arch Ophthalmol 72: 346-350.
- Whitcomb MP, Jeffries CD, Weise RW (1981) Curvularia lunata in experimental phaeohyphomycosis. Mycopathologia 75(2): 81-88.
- Tripathi P, Kukreja N, Singh BP, Arora N (2009) Serine protease activity of Cur l 1 from Curvularia lunata augments Th2 response in mice. J Clin Immunol 29(3): 292-302.
- Figueiredo-Carvalho MHG, Ramos LS, Barbedo LS, Jean Carlos Almeida de Oliveira, André Luis Souza dos Santos, et al. (2017) Relationship between the Antifungal Susceptibility Profile and the Production of Virulence-Related Hydrolytic Enzymes in Brazilian Clinical Strains of Candida glabrata. Mediators Inflamm, pp. 8952878.
- Madhu SN, Jha KK, Karthyayani AP, Gajjar DU (2018) Ex vivo caprine model to study virulence factors in keratitis. J Opthalmic Vis Res 13(4): 383-391.




